
- Customer Reviews
- Extended Essays
- IB Internal Assessment
- Theory of Knowledge
- Literature Review
- Dissertations
- Essay Writing
- Research Writing
- Assignment Help
- Capstone Projects
- College Application
- Online Class

Abstract for Research Proposal: Types and How to Write It
by Antony W
June 26, 2024
An abstract in a research proposal summarizes the main aspect of the assignment in a given sequence in 300 words or less. It highlights the purpose of the study, the research problem, design of the study, findings, summary of your interpretations and conclusions.
For what it’s worth, the abstract of your research proposal should give a clear and concise elaboration of the major aspects of an issue you’ve investigated.
In this guide, you’ll learn how to write an abstract for any research proposal. We’ll look at why an abstract is important, the types of abstracts, writing style, and what to avoid when it comes to writing an abstract for your research proposal.
Types of Abstracts for a Research Proposal
There are four types of abstracts that you can write for a research proposal :
- Critical abstract
- Descriptive abstract
- Informative abstract
- Highlight abstract
1. Critical abstract
A critical abstract in a research proposal describes the primary findings and gives a solid judgment on the validity, completeness, and reliability of the study. It’s your responsibility as a researcher to evaluate your work and then compare it with already existing work on the same subject.
Because a critical abstract includes an additional commentary, it tends to longer. Often, the length falls between 400 and 500 words. However, do keep in mind that this type of an abstract is very are, which means your instructor may never ask you to write a critical abstract for your research proposal.
2. Highlight Abstract
A highlight abstract is a piece of writing that can’t stand independent of its associated document. It uses incomplete and leading remarks, with the primary goal of grabbing the attention of the reader to the study.
Professors have made it clear that a highlight abstract is not by itself a true abstract to use in a research proposal. Since it cannot stand on its away separate from the associated article, it’s unlikely that your teacher will ask you to use it in academic writing.
3. Descriptive abstract
A descriptive abstract gives a short description of the research proposal. It may include purpose, method, and the scope of the research, and it’s often 100 words or less in length. Some people consider it to be an outline of the research proposal rather than an actual abstract for the document.
While a descriptive abstract describes the type of information a reader will find in a research proposal, it neither critics the work nor provides results and conclusion of the study.
4. Informative Abstract
Many abstracts in academic writing are informative. They don’t analyze the study or investigation that you propose, but they explain a research project in a way that they can stand independently. In other words, an informative abstract gives an explanation for the main arguments, evidence, and significant results.
In addition to featuring purpose, method, and scope, an informative abstract also include the results, conclusion, as well as the recommendation of the author. As for the length, an informative abstract should not be more than 300 words.
How to Write an Abstract for a Research Proposal
Of the four type of abstracts that we’ve discussed above, an informative abstract is what you’ll need to write in your research proposal. Writing an abstract for a research proposal isn’t difficult at all. You only need to know what to write and how to write it, and you’re good to get started.
1. Write in Active Voice
First, use active voice when writing an abstract for your research proposal. However, this doesn’t mean you should avoid passive voice in entirety. If you find that some sentences can’t make sense unless with passive sentence construction, feel free to bend this rule somewhat.
Second, make sure your sentences are concise and complete. Refrain from using ambiguous words. Keep the language simple instead.
Lastly, never use present or future tense to write an abstract for a research proposal. You’re reporting a study that you’ve already conducted and therefore writing in past sense makes the most sense.
Your abstract should come immediately after the title page. Write in block format without paragraph indentations. The abstract should not be more than 300 words long and the page should not have a number. The word “Abstract” in your research proposal should be center aligned in the page, unless otherwise stated.
In addition to these formatting rules, the last sentence of your abstract should summarize the application to practice or the conclusions of your study. In the case where it seems appropriate, you might want follow this by statement that suggests a need for additional research.
3. Time to Write the Abstract
There are no hard rules on when to write an abstract for a research proposal. Some students choose to write the section first while others choose to write it last. We strongly recommend that you write the abstract last because it’s a summary of the whole paper. You can also write it in the beginning if you’ve already outlined your draft and know what you want to talk about even before you start writing.
Your informative abstract is subject to frequent changes as you work on your paper, and that holds whether you write the section first or last. Be flexible and tweak this part of the assignment as necessary. Also, make sure you report statistical findings in parentheses.
Read abstract to be sure the summary of the study agrees with what you’ve written in your proposal. As we mentioned earlier, this section is subject to change depending on the direction your research takes. So make sure you identify and correct any anomalies if any.
Mistakes to Avoid When Writing an Abstract for Research Proposal
To wind up this guide, here are some of the most common mistakes that you should avoid when writing an abstract for your research proposal:
- Avoid giving a lengthy background
- Don’t include citations to other people’s work
- An abstract shouldn’t include a table, figure, image, or any kind of illustration
- Don’t include terms that are difficult to understand
About the author
Antony W is a professional writer and coach at Help for Assessment. He spends countless hours every day researching and writing great content filled with expert advice on how to write engaging essays, research papers, and assignments.
- Features for Creative Writers
- Features for Work
- Features for Higher Education
- Features for Teachers
- Features for Non-Native Speakers
- Learn Blog Grammar Guide Community Events FAQ
- Grammar Guide
How to Write an Abstract (With Examples)

Sarah Oakley

Table of Contents
What is an abstract in a paper, how long should an abstract be, 5 steps for writing an abstract, examples of an abstract, how prowritingaid can help you write an abstract.
If you are writing a scientific research paper or a book proposal, you need to know how to write an abstract, which summarizes the contents of the paper or book.
When researchers are looking for peer-reviewed papers to use in their studies, the first place they will check is the abstract to see if it applies to their work. Therefore, your abstract is one of the most important parts of your entire paper.
In this article, we’ll explain what an abstract is, what it should include, and how to write one.
An abstract is a concise summary of the details within a report. Some abstracts give more details than others, but the main things you’ll be talking about are why you conducted the research, what you did, and what the results show.
When a reader is deciding whether to read your paper completely, they will first look at the abstract. You need to be concise in your abstract and give the reader the most important information so they can determine if they want to read the whole paper.
Remember that an abstract is the last thing you’ll want to write for the research paper because it directly references parts of the report. If you haven’t written the report, you won’t know what to include in your abstract.
If you are writing a paper for a journal or an assignment, the publication or academic institution might have specific formatting rules for how long your abstract should be. However, if they don’t, most abstracts are between 150 and 300 words long.
A short word count means your writing has to be precise and without filler words or phrases. Once you’ve written a first draft, you can always use an editing tool, such as ProWritingAid, to identify areas where you can reduce words and increase readability.
If your abstract is over the word limit, and you’ve edited it but still can’t figure out how to reduce it further, your abstract might include some things that aren’t needed. Here’s a list of three elements you can remove from your abstract:
Discussion : You don’t need to go into detail about the findings of your research because your reader will find your discussion within the paper.
Definition of terms : Your readers are interested the field you are writing about, so they are likely to understand the terms you are using. If not, they can always look them up. Your readers do not expect you to give a definition of terms in your abstract.
References and citations : You can mention there have been studies that support or have inspired your research, but you do not need to give details as the reader will find them in your bibliography.

Good writing = better grades
ProWritingAid will help you improve the style, strength, and clarity of all your assignments.
If you’ve never written an abstract before, and you’re wondering how to write an abstract, we’ve got some steps for you to follow. It’s best to start with planning your abstract, so we’ve outlined the details you need to include in your plan before you write.
Remember to consider your audience when you’re planning and writing your abstract. They are likely to skim read your abstract, so you want to be sure your abstract delivers all the information they’re expecting to see at key points.
1. What Should an Abstract Include?
Abstracts have a lot of information to cover in a short number of words, so it’s important to know what to include. There are three elements that need to be present in your abstract:
Your context is the background for where your research sits within your field of study. You should briefly mention any previous scientific papers or experiments that have led to your hypothesis and how research develops in those studies.
Your hypothesis is your prediction of what your study will show. As you are writing your abstract after you have conducted your research, you should still include your hypothesis in your abstract because it shows the motivation for your paper.
Throughout your abstract, you also need to include keywords and phrases that will help researchers to find your article in the databases they’re searching. Make sure the keywords are specific to your field of study and the subject you’re reporting on, otherwise your article might not reach the relevant audience.
2. Can You Use First Person in an Abstract?
You might think that first person is too informal for a research paper, but it’s not. Historically, writers of academic reports avoided writing in first person to uphold the formality standards of the time. However, first person is more accepted in research papers in modern times.
If you’re still unsure whether to write in first person for your abstract, refer to any style guide rules imposed by the journal you’re writing for or your teachers if you are writing an assignment.
3. Abstract Structure
Some scientific journals have strict rules on how to structure an abstract, so it’s best to check those first. If you don’t have any style rules to follow, try using the IMRaD structure, which stands for Introduction, Methodology, Results, and Discussion.

Following the IMRaD structure, start with an introduction. The amount of background information you should include depends on your specific research area. Adding a broad overview gives you less room to include other details. Remember to include your hypothesis in this section.
The next part of your abstract should cover your methodology. Try to include the following details if they apply to your study:
What type of research was conducted?
How were the test subjects sampled?
What were the sample sizes?
What was done to each group?
How long was the experiment?
How was data recorded and interpreted?
Following the methodology, include a sentence or two about the results, which is where your reader will determine if your research supports or contradicts their own investigations.
The results are also where most people will want to find out what your outcomes were, even if they are just mildly interested in your research area. You should be specific about all the details but as concise as possible.
The last few sentences are your conclusion. It needs to explain how your findings affect the context and whether your hypothesis was correct. Include the primary take-home message, additional findings of importance, and perspective. Also explain whether there is scope for further research into the subject of your report.
Your conclusion should be honest and give the reader the ultimate message that your research shows. Readers trust the conclusion, so make sure you’re not fabricating the results of your research. Some readers won’t read your entire paper, but this section will tell them if it’s worth them referencing it in their own study.
4. How to Start an Abstract
The first line of your abstract should give your reader the context of your report by providing background information. You can use this sentence to imply the motivation for your research.
You don’t need to use a hook phrase or device in your first sentence to grab the reader’s attention. Your reader will look to establish relevance quickly, so readability and clarity are more important than trying to persuade the reader to read on.
5. How to Format an Abstract
Most abstracts use the same formatting rules, which help the reader identify the abstract so they know where to look for it.
Here’s a list of formatting guidelines for writing an abstract:
Stick to one paragraph
Use block formatting with no indentation at the beginning
Put your abstract straight after the title and acknowledgements pages
Use present or past tense, not future tense
There are two primary types of abstract you could write for your paper—descriptive and informative.
An informative abstract is the most common, and they follow the structure mentioned previously. They are longer than descriptive abstracts because they cover more details.
Descriptive abstracts differ from informative abstracts, as they don’t include as much discussion or detail. The word count for a descriptive abstract is between 50 and 150 words.
Here is an example of an informative abstract:
A growing trend exists for authors to employ a more informal writing style that uses “we” in academic writing to acknowledge one’s stance and engagement. However, few studies have compared the ways in which the first-person pronoun “we” is used in the abstracts and conclusions of empirical papers. To address this lacuna in the literature, this study conducted a systematic corpus analysis of the use of “we” in the abstracts and conclusions of 400 articles collected from eight leading electrical and electronic (EE) engineering journals. The abstracts and conclusions were extracted to form two subcorpora, and an integrated framework was applied to analyze and seek to explain how we-clusters and we-collocations were employed. Results revealed whether authors’ use of first-person pronouns partially depends on a journal policy. The trend of using “we” showed that a yearly increase occurred in the frequency of “we” in EE journal papers, as well as the existence of three “we-use” types in the article conclusions and abstracts: exclusive, inclusive, and ambiguous. Other possible “we-use” alternatives such as “I” and other personal pronouns were used very rarely—if at all—in either section. These findings also suggest that the present tense was used more in article abstracts, but the present perfect tense was the most preferred tense in article conclusions. Both research and pedagogical implications are proffered and critically discussed.
Wang, S., Tseng, W.-T., & Johanson, R. (2021). To We or Not to We: Corpus-Based Research on First-Person Pronoun Use in Abstracts and Conclusions. SAGE Open, 11(2).
Here is an example of a descriptive abstract:
From the 1850s to the present, considerable criminological attention has focused on the development of theoretically-significant systems for classifying crime. This article reviews and attempts to evaluate a number of these efforts, and we conclude that further work on this basic task is needed. The latter part of the article explicates a conceptual foundation for a crime pattern classification system, and offers a preliminary taxonomy of crime.
Farr, K. A., & Gibbons, D. C. (1990). Observations on the Development of Crime Categories. International Journal of Offender Therapy and Comparative Criminology, 34(3), 223–237.
If you want to ensure your abstract is grammatically correct and easy to read, you can use ProWritingAid to edit it. The software integrates with Microsoft Word, Google Docs, and most web browsers, so you can make the most of it wherever you’re writing your paper.
Before you edit with ProWritingAid, make sure the suggestions you are seeing are relevant for your document by changing the document type to “Abstract” within the Academic writing style section.
You can use the Readability report to check your abstract for places to improve the clarity of your writing. Some suggestions might show you where to remove words, which is great if you’re over your word count.
We hope the five steps and examples we’ve provided help you write a great abstract for your research paper.
Get started with ProWritingAid
Drop us a line or let's stay in touch via :
Writing an Abstract for Your Research Paper
Definition and Purpose of Abstracts
An abstract is a short summary of your (published or unpublished) research paper, usually about a paragraph (c. 6-7 sentences, 150-250 words) long. A well-written abstract serves multiple purposes:
- an abstract lets readers get the gist or essence of your paper or article quickly, in order to decide whether to read the full paper;
- an abstract prepares readers to follow the detailed information, analyses, and arguments in your full paper;
- and, later, an abstract helps readers remember key points from your paper.
It’s also worth remembering that search engines and bibliographic databases use abstracts, as well as the title, to identify key terms for indexing your published paper. So what you include in your abstract and in your title are crucial for helping other researchers find your paper or article.
If you are writing an abstract for a course paper, your professor may give you specific guidelines for what to include and how to organize your abstract. Similarly, academic journals often have specific requirements for abstracts. So in addition to following the advice on this page, you should be sure to look for and follow any guidelines from the course or journal you’re writing for.
The Contents of an Abstract
Abstracts contain most of the following kinds of information in brief form. The body of your paper will, of course, develop and explain these ideas much more fully. As you will see in the samples below, the proportion of your abstract that you devote to each kind of information—and the sequence of that information—will vary, depending on the nature and genre of the paper that you are summarizing in your abstract. And in some cases, some of this information is implied, rather than stated explicitly. The Publication Manual of the American Psychological Association , which is widely used in the social sciences, gives specific guidelines for what to include in the abstract for different kinds of papers—for empirical studies, literature reviews or meta-analyses, theoretical papers, methodological papers, and case studies.
Here are the typical kinds of information found in most abstracts:
- the context or background information for your research; the general topic under study; the specific topic of your research
- the central questions or statement of the problem your research addresses
- what’s already known about this question, what previous research has done or shown
- the main reason(s) , the exigency, the rationale , the goals for your research—Why is it important to address these questions? Are you, for example, examining a new topic? Why is that topic worth examining? Are you filling a gap in previous research? Applying new methods to take a fresh look at existing ideas or data? Resolving a dispute within the literature in your field? . . .
- your research and/or analytical methods
- your main findings , results , or arguments
- the significance or implications of your findings or arguments.
Your abstract should be intelligible on its own, without a reader’s having to read your entire paper. And in an abstract, you usually do not cite references—most of your abstract will describe what you have studied in your research and what you have found and what you argue in your paper. In the body of your paper, you will cite the specific literature that informs your research.
When to Write Your Abstract
Although you might be tempted to write your abstract first because it will appear as the very first part of your paper, it’s a good idea to wait to write your abstract until after you’ve drafted your full paper, so that you know what you’re summarizing.
What follows are some sample abstracts in published papers or articles, all written by faculty at UW-Madison who come from a variety of disciplines. We have annotated these samples to help you see the work that these authors are doing within their abstracts.
Choosing Verb Tenses within Your Abstract
The social science sample (Sample 1) below uses the present tense to describe general facts and interpretations that have been and are currently true, including the prevailing explanation for the social phenomenon under study. That abstract also uses the present tense to describe the methods, the findings, the arguments, and the implications of the findings from their new research study. The authors use the past tense to describe previous research.
The humanities sample (Sample 2) below uses the past tense to describe completed events in the past (the texts created in the pulp fiction industry in the 1970s and 80s) and uses the present tense to describe what is happening in those texts, to explain the significance or meaning of those texts, and to describe the arguments presented in the article.
The science samples (Samples 3 and 4) below use the past tense to describe what previous research studies have done and the research the authors have conducted, the methods they have followed, and what they have found. In their rationale or justification for their research (what remains to be done), they use the present tense. They also use the present tense to introduce their study (in Sample 3, “Here we report . . .”) and to explain the significance of their study (In Sample 3, This reprogramming . . . “provides a scalable cell source for. . .”).
Sample Abstract 1
From the social sciences.
Reporting new findings about the reasons for increasing economic homogamy among spouses
Gonalons-Pons, Pilar, and Christine R. Schwartz. “Trends in Economic Homogamy: Changes in Assortative Mating or the Division of Labor in Marriage?” Demography , vol. 54, no. 3, 2017, pp. 985-1005.
![abstract in research proposal sample “The growing economic resemblance of spouses has contributed to rising inequality by increasing the number of couples in which there are two high- or two low-earning partners. [Annotation for the previous sentence: The first sentence introduces the topic under study (the “economic resemblance of spouses”). This sentence also implies the question underlying this research study: what are the various causes—and the interrelationships among them—for this trend?] The dominant explanation for this trend is increased assortative mating. Previous research has primarily relied on cross-sectional data and thus has been unable to disentangle changes in assortative mating from changes in the division of spouses’ paid labor—a potentially key mechanism given the dramatic rise in wives’ labor supply. [Annotation for the previous two sentences: These next two sentences explain what previous research has demonstrated. By pointing out the limitations in the methods that were used in previous studies, they also provide a rationale for new research.] We use data from the Panel Study of Income Dynamics (PSID) to decompose the increase in the correlation between spouses’ earnings and its contribution to inequality between 1970 and 2013 into parts due to (a) changes in assortative mating, and (b) changes in the division of paid labor. [Annotation for the previous sentence: The data, research and analytical methods used in this new study.] Contrary to what has often been assumed, the rise of economic homogamy and its contribution to inequality is largely attributable to changes in the division of paid labor rather than changes in sorting on earnings or earnings potential. Our findings indicate that the rise of economic homogamy cannot be explained by hypotheses centered on meeting and matching opportunities, and they show where in this process inequality is generated and where it is not.” (p. 985) [Annotation for the previous two sentences: The major findings from and implications and significance of this study.]](https://writing.wisc.edu/wp-content/uploads/sites/535/2019/08/Abstract-1.png)
Sample Abstract 2
From the humanities.
Analyzing underground pulp fiction publications in Tanzania, this article makes an argument about the cultural significance of those publications
Emily Callaci. “Street Textuality: Socialism, Masculinity, and Urban Belonging in Tanzania’s Pulp Fiction Publishing Industry, 1975-1985.” Comparative Studies in Society and History , vol. 59, no. 1, 2017, pp. 183-210.
![abstract in research proposal sample “From the mid-1970s through the mid-1980s, a network of young urban migrant men created an underground pulp fiction publishing industry in the city of Dar es Salaam. [Annotation for the previous sentence: The first sentence introduces the context for this research and announces the topic under study.] As texts that were produced in the underground economy of a city whose trajectory was increasingly charted outside of formalized planning and investment, these novellas reveal more than their narrative content alone. These texts were active components in the urban social worlds of the young men who produced them. They reveal a mode of urbanism otherwise obscured by narratives of decolonization, in which urban belonging was constituted less by national citizenship than by the construction of social networks, economic connections, and the crafting of reputations. This article argues that pulp fiction novellas of socialist era Dar es Salaam are artifacts of emergent forms of male sociability and mobility. In printing fictional stories about urban life on pilfered paper and ink, and distributing their texts through informal channels, these writers not only described urban communities, reputations, and networks, but also actually created them.” (p. 210) [Annotation for the previous sentences: The remaining sentences in this abstract interweave other essential information for an abstract for this article. The implied research questions: What do these texts mean? What is their historical and cultural significance, produced at this time, in this location, by these authors? The argument and the significance of this analysis in microcosm: these texts “reveal a mode or urbanism otherwise obscured . . .”; and “This article argues that pulp fiction novellas. . . .” This section also implies what previous historical research has obscured. And through the details in its argumentative claims, this section of the abstract implies the kinds of methods the author has used to interpret the novellas and the concepts under study (e.g., male sociability and mobility, urban communities, reputations, network. . . ).]](https://writing.wisc.edu/wp-content/uploads/sites/535/2019/08/Abstract-2.png)
Sample Abstract/Summary 3
From the sciences.
Reporting a new method for reprogramming adult mouse fibroblasts into induced cardiac progenitor cells
Lalit, Pratik A., Max R. Salick, Daryl O. Nelson, Jayne M. Squirrell, Christina M. Shafer, Neel G. Patel, Imaan Saeed, Eric G. Schmuck, Yogananda S. Markandeya, Rachel Wong, Martin R. Lea, Kevin W. Eliceiri, Timothy A. Hacker, Wendy C. Crone, Michael Kyba, Daniel J. Garry, Ron Stewart, James A. Thomson, Karen M. Downs, Gary E. Lyons, and Timothy J. Kamp. “Lineage Reprogramming of Fibroblasts into Proliferative Induced Cardiac Progenitor Cells by Defined Factors.” Cell Stem Cell , vol. 18, 2016, pp. 354-367.
![abstract in research proposal sample “Several studies have reported reprogramming of fibroblasts into induced cardiomyocytes; however, reprogramming into proliferative induced cardiac progenitor cells (iCPCs) remains to be accomplished. [Annotation for the previous sentence: The first sentence announces the topic under study, summarizes what’s already known or been accomplished in previous research, and signals the rationale and goals are for the new research and the problem that the new research solves: How can researchers reprogram fibroblasts into iCPCs?] Here we report that a combination of 11 or 5 cardiac factors along with canonical Wnt and JAK/STAT signaling reprogrammed adult mouse cardiac, lung, and tail tip fibroblasts into iCPCs. The iCPCs were cardiac mesoderm-restricted progenitors that could be expanded extensively while maintaining multipo-tency to differentiate into cardiomyocytes, smooth muscle cells, and endothelial cells in vitro. Moreover, iCPCs injected into the cardiac crescent of mouse embryos differentiated into cardiomyocytes. iCPCs transplanted into the post-myocardial infarction mouse heart improved survival and differentiated into cardiomyocytes, smooth muscle cells, and endothelial cells. [Annotation for the previous four sentences: The methods the researchers developed to achieve their goal and a description of the results.] Lineage reprogramming of adult somatic cells into iCPCs provides a scalable cell source for drug discovery, disease modeling, and cardiac regenerative therapy.” (p. 354) [Annotation for the previous sentence: The significance or implications—for drug discovery, disease modeling, and therapy—of this reprogramming of adult somatic cells into iCPCs.]](https://writing.wisc.edu/wp-content/uploads/sites/535/2019/08/Abstract-3.png)
Sample Abstract 4, a Structured Abstract
Reporting results about the effectiveness of antibiotic therapy in managing acute bacterial sinusitis, from a rigorously controlled study
Note: This journal requires authors to organize their abstract into four specific sections, with strict word limits. Because the headings for this structured abstract are self-explanatory, we have chosen not to add annotations to this sample abstract.
Wald, Ellen R., David Nash, and Jens Eickhoff. “Effectiveness of Amoxicillin/Clavulanate Potassium in the Treatment of Acute Bacterial Sinusitis in Children.” Pediatrics , vol. 124, no. 1, 2009, pp. 9-15.
“OBJECTIVE: The role of antibiotic therapy in managing acute bacterial sinusitis (ABS) in children is controversial. The purpose of this study was to determine the effectiveness of high-dose amoxicillin/potassium clavulanate in the treatment of children diagnosed with ABS.
METHODS : This was a randomized, double-blind, placebo-controlled study. Children 1 to 10 years of age with a clinical presentation compatible with ABS were eligible for participation. Patients were stratified according to age (<6 or ≥6 years) and clinical severity and randomly assigned to receive either amoxicillin (90 mg/kg) with potassium clavulanate (6.4 mg/kg) or placebo. A symptom survey was performed on days 0, 1, 2, 3, 5, 7, 10, 20, and 30. Patients were examined on day 14. Children’s conditions were rated as cured, improved, or failed according to scoring rules.
RESULTS: Two thousand one hundred thirty-five children with respiratory complaints were screened for enrollment; 139 (6.5%) had ABS. Fifty-eight patients were enrolled, and 56 were randomly assigned. The mean age was 6630 months. Fifty (89%) patients presented with persistent symptoms, and 6 (11%) presented with nonpersistent symptoms. In 24 (43%) children, the illness was classified as mild, whereas in the remaining 32 (57%) children it was severe. Of the 28 children who received the antibiotic, 14 (50%) were cured, 4 (14%) were improved, 4(14%) experienced treatment failure, and 6 (21%) withdrew. Of the 28children who received placebo, 4 (14%) were cured, 5 (18%) improved, and 19 (68%) experienced treatment failure. Children receiving the antibiotic were more likely to be cured (50% vs 14%) and less likely to have treatment failure (14% vs 68%) than children receiving the placebo.
CONCLUSIONS : ABS is a common complication of viral upper respiratory infections. Amoxicillin/potassium clavulanate results in significantly more cures and fewer failures than placebo, according to parental report of time to resolution.” (9)
Some Excellent Advice about Writing Abstracts for Basic Science Research Papers, by Professor Adriano Aguzzi from the Institute of Neuropathology at the University of Zurich:

Academic and Professional Writing
This is an accordion element with a series of buttons that open and close related content panels.
Analysis Papers
Reading Poetry
A Short Guide to Close Reading for Literary Analysis
Using Literary Quotations
Play Reviews
Writing a Rhetorical Précis to Analyze Nonfiction Texts
Incorporating Interview Data
Grant Proposals
Planning and Writing a Grant Proposal: The Basics
Additional Resources for Grants and Proposal Writing
Job Materials and Application Essays
Writing Personal Statements for Ph.D. Programs
- Before you begin: useful tips for writing your essay
- Guided brainstorming exercises
- Get more help with your essay
- Frequently Asked Questions
Resume Writing Tips
CV Writing Tips
Cover Letters
Business Letters
Proposals and Dissertations
Resources for Proposal Writers
Resources for Dissertators
Research Papers
Planning and Writing Research Papers
Quoting and Paraphrasing
Writing Annotated Bibliographies
Creating Poster Presentations
Thank-You Notes
Advice for Students Writing Thank-You Notes to Donors
Reading for a Review
Critical Reviews
Writing a Review of Literature
Scientific Reports
Scientific Report Format
Sample Lab Assignment
Writing for the Web
Writing an Effective Blog Post
Writing for Social Media: A Guide for Academics
- Resources Home 🏠
- Try SciSpace Copilot
- Search research papers
- Add Copilot Extension
- Try AI Detector
- Try Paraphraser
- Try Citation Generator
- April Papers
- June Papers
- July Papers

Abstract Writing: A Step-by-Step Guide With Tips & Examples

Table of Contents

Introduction
Abstracts of research papers have always played an essential role in describing your research concisely and clearly to researchers and editors of journals, enticing them to continue reading. However, with the widespread availability of scientific databases, the need to write a convincing abstract is more crucial now than during the time of paper-bound manuscripts.
Abstracts serve to "sell" your research and can be compared with your "executive outline" of a resume or, rather, a formal summary of the critical aspects of your work. Also, it can be the "gist" of your study. Since most educational research is done online, it's a sign that you have a shorter time for impressing your readers, and have more competition from other abstracts that are available to be read.
The APCI (Academic Publishing and Conferences International) articulates 12 issues or points considered during the final approval process for conferences & journals and emphasises the importance of writing an abstract that checks all these boxes (12 points). Since it's the only opportunity you have to captivate your readers, you must invest time and effort in creating an abstract that accurately reflects the critical points of your research.
With that in mind, let’s head over to understand and discover the core concept and guidelines to create a substantial abstract. Also, learn how to organise the ideas or plots into an effective abstract that will be awe-inspiring to the readers you want to reach.
What is Abstract? Definition and Overview
The word "Abstract' is derived from Latin abstractus meaning "drawn off." This etymological meaning also applies to art movements as well as music, like abstract expressionism. In this context, it refers to the revealing of the artist's intention.
Based on this, you can determine the meaning of an abstract: A condensed research summary. It must be self-contained and independent of the body of the research. However, it should outline the subject, the strategies used to study the problem, and the methods implemented to attain the outcomes. The specific elements of the study differ based on the area of study; however, together, it must be a succinct summary of the entire research paper.
Abstracts are typically written at the end of the paper, even though it serves as a prologue. In general, the abstract must be in a position to:
- Describe the paper.
- Identify the problem or the issue at hand.
- Explain to the reader the research process, the results you came up with, and what conclusion you've reached using these results.
- Include keywords to guide your strategy and the content.
Furthermore, the abstract you submit should not reflect upon any of the following elements:
- Examine, analyse or defend the paper or your opinion.
- What you want to study, achieve or discover.
- Be redundant or irrelevant.
After reading an abstract, your audience should understand the reason - what the research was about in the first place, what the study has revealed and how it can be utilised or can be used to benefit others. You can understand the importance of abstract by knowing the fact that the abstract is the most frequently read portion of any research paper. In simpler terms, it should contain all the main points of the research paper.
What is the Purpose of an Abstract?
Abstracts are typically an essential requirement for research papers; however, it's not an obligation to preserve traditional reasons without any purpose. Abstracts allow readers to scan the text to determine whether it is relevant to their research or studies. The abstract allows other researchers to decide if your research paper can provide them with some additional information. A good abstract paves the interest of the audience to pore through your entire paper to find the content or context they're searching for.
Abstract writing is essential for indexing, as well. The Digital Repository of academic papers makes use of abstracts to index the entire content of academic research papers. Like meta descriptions in the regular Google outcomes, abstracts must include keywords that help researchers locate what they seek.
Types of Abstract
Informative and Descriptive are two kinds of abstracts often used in scientific writing.
A descriptive abstract gives readers an outline of the author's main points in their study. The reader can determine if they want to stick to the research work, based on their interest in the topic. An abstract that is descriptive is similar to the contents table of books, however, the format of an abstract depicts complete sentences encapsulated in one paragraph. It is unfortunate that the abstract can't be used as a substitute for reading a piece of writing because it's just an overview, which omits readers from getting an entire view. Also, it cannot be a way to fill in the gaps the reader may have after reading this kind of abstract since it does not contain crucial information needed to evaluate the article.
To conclude, a descriptive abstract is:
- A simple summary of the task, just summarises the work, but some researchers think it is much more of an outline
- Typically, the length is approximately 100 words. It is too short when compared to an informative abstract.
- A brief explanation but doesn't provide the reader with the complete information they need;
- An overview that omits conclusions and results
An informative abstract is a comprehensive outline of the research. There are times when people rely on the abstract as an information source. And the reason is why it is crucial to provide entire data of particular research. A well-written, informative abstract could be a good substitute for the remainder of the paper on its own.
A well-written abstract typically follows a particular style. The author begins by providing the identifying information, backed by citations and other identifiers of the papers. Then, the major elements are summarised to make the reader aware of the study. It is followed by the methodology and all-important findings from the study. The conclusion then presents study results and ends the abstract with a comprehensive summary.
In a nutshell, an informative abstract:
- Has a length that can vary, based on the subject, but is not longer than 300 words.
- Contains all the content-like methods and intentions
- Offers evidence and possible recommendations.
Informative Abstracts are more frequent than descriptive abstracts because of their extensive content and linkage to the topic specifically. You should select different types of abstracts to papers based on their length: informative abstracts for extended and more complex abstracts and descriptive ones for simpler and shorter research papers.
What are the Characteristics of a Good Abstract?
- A good abstract clearly defines the goals and purposes of the study.
- It should clearly describe the research methodology with a primary focus on data gathering, processing, and subsequent analysis.
- A good abstract should provide specific research findings.
- It presents the principal conclusions of the systematic study.
- It should be concise, clear, and relevant to the field of study.
- A well-designed abstract should be unifying and coherent.
- It is easy to grasp and free of technical jargon.
- It is written impartially and objectively.
What are the various sections of an ideal Abstract?
By now, you must have gained some concrete idea of the essential elements that your abstract needs to convey . Accordingly, the information is broken down into six key sections of the abstract, which include:
An Introduction or Background
Research methodology, objectives and goals, limitations.
Let's go over them in detail.
The introduction, also known as background, is the most concise part of your abstract. Ideally, it comprises a couple of sentences. Some researchers only write one sentence to introduce their abstract. The idea behind this is to guide readers through the key factors that led to your study.
It's understandable that this information might seem difficult to explain in a couple of sentences. For example, think about the following two questions like the background of your study:
- What is currently available about the subject with respect to the paper being discussed?
- What isn't understood about this issue? (This is the subject of your research)
While writing the abstract’s introduction, make sure that it is not lengthy. Because if it crosses the word limit, it may eat up the words meant to be used for providing other key information.
Research methodology is where you describe the theories and techniques you used in your research. It is recommended that you describe what you have done and the method you used to get your thorough investigation results. Certainly, it is the second-longest paragraph in the abstract.
In the research methodology section, it is essential to mention the kind of research you conducted; for instance, qualitative research or quantitative research (this will guide your research methodology too) . If you've conducted quantitative research, your abstract should contain information like the sample size, data collection method, sampling techniques, and duration of the study. Likewise, your abstract should reflect observational data, opinions, questionnaires (especially the non-numerical data) if you work on qualitative research.
The research objectives and goals speak about what you intend to accomplish with your research. The majority of research projects focus on the long-term effects of a project, and the goals focus on the immediate, short-term outcomes of the research. It is possible to summarise both in just multiple sentences.
In stating your objectives and goals, you give readers a picture of the scope of the study, its depth and the direction your research ultimately follows. Your readers can evaluate the results of your research against the goals and stated objectives to determine if you have achieved the goal of your research.
In the end, your readers are more attracted by the results you've obtained through your study. Therefore, you must take the time to explain each relevant result and explain how they impact your research. The results section exists as the longest in your abstract, and nothing should diminish its reach or quality.
One of the most important things you should adhere to is to spell out details and figures on the results of your research.
Instead of making a vague assertion such as, "We noticed that response rates varied greatly between respondents with high incomes and those with low incomes", Try these: "The response rate was higher for high-income respondents than those with lower incomes (59 30 percent vs. 30 percent in both cases; P<0.01)."
You're likely to encounter certain obstacles during your research. It could have been during data collection or even during conducting the sample . Whatever the issue, it's essential to inform your readers about them and their effects on the research.
Research limitations offer an opportunity to suggest further and deep research. If, for instance, you were forced to change for convenient sampling and snowball samples because of difficulties in reaching well-suited research participants, then you should mention this reason when you write your research abstract. In addition, a lack of prior studies on the subject could hinder your research.
Your conclusion should include the same number of sentences to wrap the abstract as the introduction. The majority of researchers offer an idea of the consequences of their research in this case.
Your conclusion should include three essential components:
- A significant take-home message.
- Corresponding important findings.
- The Interpretation.
Even though the conclusion of your abstract needs to be brief, it can have an enormous influence on the way that readers view your research. Therefore, make use of this section to reinforce the central message from your research. Be sure that your statements reflect the actual results and the methods you used to conduct your research.
Good Abstract Examples
Abstract example #1.
Children’s consumption behavior in response to food product placements in movies.
The abstract:
"Almost all research into the effects of brand placements on children has focused on the brand's attitudes or behavior intentions. Based on the significant differences between attitudes and behavioral intentions on one hand and actual behavior on the other hand, this study examines the impact of placements by brands on children's eating habits. Children aged 6-14 years old were shown an excerpt from the popular film Alvin and the Chipmunks and were shown places for the item Cheese Balls. Three different versions were developed with no placements, one with moderately frequent placements and the third with the highest frequency of placement. The results revealed that exposure to high-frequency places had a profound effect on snack consumption, however, there was no impact on consumer attitudes towards brands or products. The effects were not dependent on the age of the children. These findings are of major importance to researchers studying consumer behavior as well as nutrition experts as well as policy regulators."
Abstract Example #2
Social comparisons on social media: The impact of Facebook on young women’s body image concerns and mood. The abstract:
"The research conducted in this study investigated the effects of Facebook use on women's moods and body image if the effects are different from an internet-based fashion journal and if the appearance comparison tendencies moderate one or more of these effects. Participants who were female ( N = 112) were randomly allocated to spend 10 minutes exploring their Facebook account or a magazine's website or an appearance neutral control website prior to completing state assessments of body dissatisfaction, mood, and differences in appearance (weight-related and facial hair, face, and skin). Participants also completed a test of the tendency to compare appearances. The participants who used Facebook were reported to be more depressed than those who stayed on the control site. In addition, women who have the tendency to compare appearances reported more facial, hair and skin-related issues following Facebook exposure than when they were exposed to the control site. Due to its popularity it is imperative to conduct more research to understand the effect that Facebook affects the way people view themselves."
Abstract Example #3
The Relationship Between Cell Phone Use and Academic Performance in a Sample of U.S. College Students
"The cellphone is always present on campuses of colleges and is often utilised in situations in which learning takes place. The study examined the connection between the use of cell phones and the actual grades point average (GPA) after adjusting for predictors that are known to be a factor. In the end 536 students in the undergraduate program from 82 self-reported majors of an enormous, public institution were studied. Hierarchical analysis ( R 2 = .449) showed that use of mobile phones is significantly ( p < .001) and negative (b equal to -.164) connected to the actual college GPA, after taking into account factors such as demographics, self-efficacy in self-regulated learning, self-efficacy to improve academic performance, and the actual high school GPA that were all important predictors ( p < .05). Therefore, after adjusting for other known predictors increasing cell phone usage was associated with lower academic performance. While more research is required to determine the mechanisms behind these results, they suggest the need to educate teachers and students to the possible academic risks that are associated with high-frequency mobile phone usage."
Quick tips on writing a good abstract
There exists a common dilemma among early age researchers whether to write the abstract at first or last? However, it's recommended to compose your abstract when you've completed the research since you'll have all the information to give to your readers. You can, however, write a draft at the beginning of your research and add in any gaps later.
If you find abstract writing a herculean task, here are the few tips to help you with it:
1. Always develop a framework to support your abstract
Before writing, ensure you create a clear outline for your abstract. Divide it into sections and draw the primary and supporting elements in each one. You can include keywords and a few sentences that convey the essence of your message.
2. Review Other Abstracts
Abstracts are among the most frequently used research documents, and thousands of them were written in the past. Therefore, prior to writing yours, take a look at some examples from other abstracts. There are plenty of examples of abstracts for dissertations in the dissertation and thesis databases.
3. Avoid Jargon To the Maximum
When you write your abstract, focus on simplicity over formality. You should write in simple language, and avoid excessive filler words or ambiguous sentences. Keep in mind that your abstract must be readable to those who aren't acquainted with your subject.
4. Focus on Your Research
It's a given fact that the abstract you write should be about your research and the findings you've made. It is not the right time to mention secondary and primary data sources unless it's absolutely required.
Conclusion: How to Structure an Interesting Abstract?
Abstracts are a short outline of your essay. However, it's among the most important, if not the most important. The process of writing an abstract is not straightforward. A few early-age researchers tend to begin by writing it, thinking they are doing it to "tease" the next step (the document itself). However, it is better to treat it as a spoiler.
The simple, concise style of the abstract lends itself to a well-written and well-investigated study. If your research paper doesn't provide definitive results, or the goal of your research is questioned, so will the abstract. Thus, only write your abstract after witnessing your findings and put your findings in the context of a larger scenario.
The process of writing an abstract can be daunting, but with these guidelines, you will succeed. The most efficient method of writing an excellent abstract is to centre the primary points of your abstract, including the research question and goals methods, as well as key results.
Interested in learning more about dedicated research solutions? Go to the SciSpace product page to find out how our suite of products can help you simplify your research workflows so you can focus on advancing science.

The best-in-class solution is equipped with features such as literature search and discovery, profile management, research writing and formatting, and so much more.
But before you go,
You might also like.

Consensus GPT vs. SciSpace GPT: Choose the Best GPT for Research

Literature Review and Theoretical Framework: Understanding the Differences

Types of Essays in Academic Writing - Quick Guide (2024)
How to Write an Abstract APA Format
Saul Mcleod, PhD
Editor-in-Chief for Simply Psychology
BSc (Hons) Psychology, MRes, PhD, University of Manchester
Saul Mcleod, PhD., is a qualified psychology teacher with over 18 years of experience in further and higher education. He has been published in peer-reviewed journals, including the Journal of Clinical Psychology.
Learn about our Editorial Process
Olivia Guy-Evans, MSc
Associate Editor for Simply Psychology
BSc (Hons) Psychology, MSc Psychology of Education
Olivia Guy-Evans is a writer and associate editor for Simply Psychology. She has previously worked in healthcare and educational sectors.
An APA abstract is a brief, comprehensive summary of the contents of an article, research paper, dissertation, or report.
It is written in accordance with the guidelines of the American Psychological Association (APA), which is a widely used format in social and behavioral sciences.
An APA abstract summarizes, usually in one paragraph of between 150–250 words, the major aspects of a research paper or dissertation in a prescribed sequence that includes:
- The rationale: the overall purpose of the study, providing a clear context for the research undertaken.
- Information regarding the method and participants: including materials/instruments, design, procedure, and data analysis.
- Main findings or trends: effectively highlighting the key outcomes of the hypotheses.
- Interpretations and conclusion(s): solidify the implications of the research.
- Keywords related to the study: assist the paper’s discoverability in academic databases.
The abstract should stand alone, be “self-contained,” and make sense to the reader in isolation from the main article.
The purpose of the abstract is to give the reader a quick overview of the essential information before reading the entire article. The abstract is placed on its own page, directly after the title page and before the main body of the paper.
Although the abstract will appear as the very first part of your paper, it’s good practice to write your abstract after you’ve drafted your full paper, so that you know what you’re summarizing.
Note : This page reflects the latest version of the APA Publication Manual (i.e., APA 7), released in October 2019.
Structure of the Abstract
[NOTE: DO NOT separate the components of the abstract – it should be written as a single paragraph. This section is separated to illustrate the abstract’s structure.]
1) The Rationale
One or two sentences describing the overall purpose of the study and the research problem(s) you investigated. You are basically justifying why this study was conducted.
- What is the importance of the research?
- Why would a reader be interested in the larger work?
- For example, are you filling a gap in previous research or applying new methods to take a fresh look at existing ideas or data?
- Women who are diagnosed with breast cancer can experience an array of psychosocial difficulties; however, social support, particularly from a spouse, has been shown to have a protective function during this time. This study examined the ways in which a woman’s daily mood, pain, and fatigue, and her spouse’s marital satisfaction predict the woman’s report of partner support in the context of breast cancer.
- The current nursing shortage, high hospital nurse job dissatisfaction, and reports of uneven quality of hospital care are not uniquely American phenomena.
- Students with special educational needs and disabilities (SEND) are more likely to exhibit behavioral difficulties than their typically developing peers. The aim of this study was to identify specific risk factors that influence variability in behavior difficulties among individuals with SEND.
2) The Method
Information regarding the participants (number, and population). One or two sentences outlining the method, explaining what was done and how. The method is described in the present tense.
- Pretest data from a larger intervention study and multilevel modeling were used to examine the effects of women’s daily mood, pain, and fatigue and average levels of mood, pain, and fatigue on women’s report of social support received from her partner, as well as how the effects of mood interacted with partners’ marital satisfaction.
- This paper presents reports from 43,000 nurses from more than 700 hospitals in the United States, Canada, England, Scotland, and Germany in 1998–1999.
- The study sample comprised 4,228 students with SEND, aged 5–15, drawn from 305 primary and secondary schools across England. Explanatory variables were measured at the individual and school levels at baseline, along with a teacher-reported measure of behavior difficulties (assessed at baseline and the 18-month follow-up).
3) The Results
One or two sentences indicating the main findings or trends found as a result of your analysis. The results are described in the present or past tense.
- Results show that on days in which women reported higher levels of negative or positive mood, as well as on days they reported more pain and fatigue, they reported receiving more support. Women who, on average, reported higher levels of positive mood tended to report receiving more support than those who, on average, reported lower positive mood. However, average levels of negative mood were not associated with support. Higher average levels of fatigue but not pain were associated with higher support. Finally, women whose husbands reported higher levels of marital satisfaction reported receiving more partner support, but husbands’ marital satisfaction did not moderate the effect of women’s mood on support.
- Nurses in countries with distinctly different healthcare systems report similar shortcomings in their work environments and the quality of hospital care. While the competence of and relation between nurses and physicians appear satisfactory, core problems in work design and workforce management threaten the provision of care.
- Hierarchical linear modeling of data revealed that differences between schools accounted for between 13% (secondary) and 15.4% (primary) of the total variance in the development of students’ behavior difficulties, with the remainder attributable to individual differences. Statistically significant risk markers for these problems across both phases of education were being male, eligibility for free school meals, being identified as a bully, and lower academic achievement. Additional risk markers specific to each phase of education at the individual and school levels are also acknowledged.
4) The Conclusion / Implications
A brief summary of your conclusions and implications of the results, described in the present tense. Explain the results and why the study is important to the reader.
- For example, what changes should be implemented as a result of the findings of the work?
- How does this work add to the body of knowledge on the topic?
Implications of these findings are discussed relative to assisting couples during this difficult time in their lives.
- Resolving these issues, which are amenable to managerial intervention, is essential to preserving patient safety and care of consistently high quality.
- Behavior difficulties are affected by risks across multiple ecological levels. Addressing any one of these potential influences is therefore likely to contribute to the reduction in the problems displayed.
The above examples of abstracts are from the following papers:
Aiken, L. H., Clarke, S. P., Sloane, D. M., Sochalski, J. A., Busse, R., Clarke, H., … & Shamian, J. (2001). Nurses’ reports on hospital care in five countries . Health affairs, 20(3) , 43-53.
Boeding, S. E., Pukay-Martin, N. D., Baucom, D. H., Porter, L. S., Kirby, J. S., Gremore, T. M., & Keefe, F. J. (2014). Couples and breast cancer: Women’s mood and partners’ marital satisfaction predicting support perception . Journal of Family Psychology, 28(5) , 675.
Oldfield, J., Humphrey, N., & Hebron, J. (2017). Risk factors in the development of behavior difficulties among students with special educational needs and disabilities: A multilevel analysis . British journal of educational psychology, 87(2) , 146-169.
5) Keywords
APA style suggests including a list of keywords at the end of the abstract. This is particularly common in academic articles and helps other researchers find your work in databases.
Keywords in an abstract should be selected to help other researchers find your work when searching an online database. These keywords should effectively represent the main topics of your study. Here are some tips for choosing keywords:
Core Concepts: Identify the most important ideas or concepts in your paper. These often include your main research topic, the methods you’ve used, or the theories you’re discussing.
Specificity: Your keywords should be specific to your research. For example, suppose your paper is about the effects of climate change on bird migration patterns in a specific region. In that case, your keywords might include “climate change,” “bird migration,” and the region’s name.
Consistency with Paper: Make sure your keywords are consistent with the terms you’ve used in your paper. For example, if you use the term “adolescent” rather than “teen” in your paper, choose “adolescent” as your keyword, not “teen.”
Jargon and Acronyms: Avoid using too much-specialized jargon or acronyms in your keywords, as these might not be understood or used by all researchers in your field.
Synonyms: Consider including synonyms of your keywords to capture as many relevant searches as possible. For example, if your paper discusses “post-traumatic stress disorder,” you might include “PTSD” as a keyword.
Remember, keywords are a tool for others to find your work, so think about what terms other researchers might use when searching for papers on your topic.
The Abstract SHOULD NOT contain:
Lengthy background or contextual information: The abstract should focus on your research and findings, not general topic background.
Undefined jargon, abbreviations, or acronyms: The abstract should be accessible to a wide audience, so avoid highly specialized terms without defining them.
Citations: Abstracts typically do not include citations, as they summarize original research.
Incomplete sentences or bulleted lists: The abstract should be a single, coherent paragraph written in complete sentences.
New information not covered in the paper: The abstract should only summarize the paper’s content.
Subjective comments or value judgments: Stick to objective descriptions of your research.
Excessive details on methods or procedures: Keep descriptions of methods brief and focused on main steps.
Speculative or inconclusive statements: The abstract should state the research’s clear findings, not hypotheses or possible interpretations.
- Any illustration, figure, table, or references to them . All visual aids, data, or extensive details should be included in the main body of your paper, not in the abstract.
- Elliptical or incomplete sentences should be avoided in an abstract . The use of ellipses (…), which could indicate incomplete thoughts or omitted text, is not appropriate in an abstract.
APA Style for Abstracts
An APA abstract must be formatted as follows:
Include the running head aligned to the left at the top of the page (professional papers only) and page number. Note, student papers do not require a running head. On the first line, center the heading “Abstract” and bold (do not underlined or italicize). Do not indent the single abstract paragraph (which begins one line below the section title). Double-space the text. Use Times New Roman font in 12 pt. Set one-inch (or 2.54 cm) margins. If you include a “keywords” section at the end of the abstract, indent the first line and italicize the word “Keywords” while leaving the keywords themselves without any formatting.
Example APA Abstract Page
Download this example as a PDF

Further Information
- APA 7th Edition Abstract and Keywords Guide
- Example APA Abstract
- How to Write a Good Abstract for a Scientific Paper or Conference Presentation
- How to Write a Lab Report
- Writing an APA paper
How long should an APA abstract be?
An APA abstract should typically be between 150 to 250 words long. However, the exact length may vary depending on specific publication or assignment guidelines. It is crucial that it succinctly summarizes the essential elements of the work, including purpose, methods, findings, and conclusions.
Where does the abstract go in an APA paper?
In an APA formatted paper, the abstract is placed on its own page, directly after the title page and before the main body of the paper. It’s typically the second page of the document. It starts with the word “Abstract” (centered and not in bold) at the top of the page, followed by the text of the abstract itself.
What are the 4 C’s of abstract writing?
The 4 C’s of abstract writing are an approach to help you create a well-structured and informative abstract. They are:
Conciseness: An abstract should briefly summarize the key points of your study. Stick to the word limit (typically between 150-250 words for an APA abstract) and avoid unnecessary details.
Clarity: Your abstract should be easy to understand. Avoid jargon and complex sentences. Clearly explain the purpose, methods, results, and conclusions of your study.
Completeness: Even though it’s brief, the abstract should provide a complete overview of your study, including the purpose, methods, key findings, and your interpretation of the results.
Cohesion: The abstract should flow logically from one point to the next, maintaining a coherent narrative about your study. It’s not just a list of disjointed elements; it’s a brief story of your research from start to finish.
What is the abstract of a psychology paper?
An abstract in a psychology paper serves as a snapshot of the paper, allowing readers to quickly understand the purpose, methodology, results, and implications of the research without reading the entire paper. It is generally between 150-250 words long.
Related Articles

Student Resources
How To Cite A YouTube Video In APA Style – With Examples

APA References Page Formatting and Example

APA Title Page (Cover Page) Format, Example, & Templates

How do I Cite a Source with Multiple Authors in APA Style?
How to Write a Psychology Essay

Lab Report Format: Step-by-Step Guide & Examples
- Affiliate Program

- UNITED STATES
- 台灣 (TAIWAN)
- TÜRKIYE (TURKEY)
- Academic Editing Services
- - Research Paper
- - Journal Manuscript
- - Dissertation
- - College & University Assignments
- Admissions Editing Services
- - Application Essay
- - Personal Statement
- - Recommendation Letter
- - Cover Letter
- - CV/Resume
- Business Editing Services
- - Business Documents
- - Report & Brochure
- - Website & Blog
- Writer Editing Services
- - Script & Screenplay
- Our Editors
- Client Reviews
- Editing & Proofreading Prices
- Wordvice Points
- Partner Discount
- Plagiarism Checker
- APA Citation Generator
- MLA Citation Generator
- Chicago Citation Generator
- Vancouver Citation Generator
- - APA Style
- - MLA Style
- - Chicago Style
- - Vancouver Style
- Writing & Editing Guide
- Academic Resources
- Admissions Resources
How to Write an Abstract for a Research Paper | Examples
What is a research paper abstract?
Research paper abstracts summarize your study quickly and succinctly to journal editors and researchers and prompt them to read further. But with the ubiquity of online publication databases, writing a compelling abstract is even more important today than it was in the days of bound paper manuscripts.
Abstracts exist to “sell” your work, and they could thus be compared to the “executive summary” of a business resume: an official briefing on what is most important about your research. Or the “gist” of your research. With the majority of academic transactions being conducted online, this means that you have even less time to impress readers–and increased competition in terms of other abstracts out there to read.
The APCI (Academic Publishing and Conferences International) notes that there are 12 questions or “points” considered in the selection process for journals and conferences and stresses the importance of having an abstract that ticks all of these boxes. Because it is often the ONLY chance you have to convince readers to keep reading, it is important that you spend time and energy crafting an abstract that faithfully represents the central parts of your study and captivates your audience.
With that in mind, follow these suggestions when structuring and writing your abstract, and learn how exactly to put these ideas into a solid abstract that will captivate your target readers.
Before Writing Your Abstract
How long should an abstract be.
All abstracts are written with the same essential objective: to give a summary of your study. But there are two basic styles of abstract: descriptive and informative . Here is a brief delineation of the two:
| Around 100-200 words (or shorter) in length; indicates the type of information found in the paper; briefly explains the background, purpose, and objective of the paper but omits the results, often the methods, and sometimes also the conclusion | |
| One paragraph to one page in length; a truncated version of your paper that summarizes every aspect of the study, including the results; acts as a “surrogate” for the research itself, standing in for the larger paper |
Of the two types of abstracts, informative abstracts are much more common, and they are widely used for submission to journals and conferences. Informative abstracts apply to lengthier and more technical research and are common in the sciences, engineering, and psychology, while descriptive abstracts are more likely used in humanities and social science papers. The best method of determining which abstract type you need to use is to follow the instructions for journal submissions and to read as many other published articles in those journals as possible.
Research Abstract Guidelines and Requirements
As any article about research writing will tell you, authors must always closely follow the specific guidelines and requirements indicated in the Guide for Authors section of their target journal’s website. The same kind of adherence to conventions should be applied to journal publications, for consideration at a conference, and even when completing a class assignment.
Each publisher has particular demands when it comes to formatting and structure. Here are some common questions addressed in the journal guidelines:
- Is there a maximum or minimum word/character length?
- What are the style and formatting requirements?
- What is the appropriate abstract type?
- Are there any specific content or organization rules that apply?
There are of course other rules to consider when composing a research paper abstract. But if you follow the stated rules the first time you submit your manuscript, you can avoid your work being thrown in the “circular file” right off the bat.
Identify Your Target Readership
The main purpose of your abstract is to lead researchers to the full text of your research paper. In scientific journals, abstracts let readers decide whether the research discussed is relevant to their own interests or study. Abstracts also help readers understand your main argument quickly. Consider these questions as you write your abstract:
- Are other academics in your field the main target of your study?
- Will your study perhaps be useful to members of the general public?
- Do your study results include the wider implications presented in the abstract?
Outlining and Writing Your Abstract
What to include in an abstract.
Just as your research paper title should cover as much ground as possible in a few short words, your abstract must cover all parts of your study in order to fully explain your paper and research. Because it must accomplish this task in the space of only a few hundred words, it is important not to include ambiguous references or phrases that will confuse the reader or mislead them about the content and objectives of your research. Follow these dos and don’ts when it comes to what kind of writing to include:
- Avoid acronyms or abbreviations since these will need to be explained in order to make sense to the reader, which takes up valuable abstract space. Instead, explain these terms in the Introduction section of the main text.
- Only use references to people or other works if they are well-known. Otherwise, avoid referencing anything outside of your study in the abstract.
- Never include tables, figures, sources, or long quotations in your abstract; you will have plenty of time to present and refer to these in the body of your paper.
Use keywords in your abstract to focus your topic
A vital search tool is the research paper keywords section, which lists the most relevant terms directly underneath the abstract. Think of these keywords as the “tubes” that readers will seek and enter—via queries on databases and search engines—to ultimately land at their destination, which is your paper. Your abstract keywords should thus be words that are commonly used in searches but should also be highly relevant to your work and found in the text of your abstract. Include 5 to 10 important words or short phrases central to your research in both the abstract and the keywords section.
For example, if you are writing a paper on the prevalence of obesity among lower classes that crosses international boundaries, you should include terms like “obesity,” “prevalence,” “international,” “lower classes,” and “cross-cultural.” These are terms that should net a wide array of people interested in your topic of study. Look at our nine rules for choosing keywords for your research paper if you need more input on this.
Research Paper Abstract Structure
As mentioned above, the abstract (especially the informative abstract) acts as a surrogate or synopsis of your research paper, doing almost as much work as the thousands of words that follow it in the body of the main text. In the hard sciences and most social sciences, the abstract includes the following sections and organizational schema.
Each section is quite compact—only a single sentence or two, although there is room for expansion if one element or statement is particularly interesting or compelling. As the abstract is almost always one long paragraph, the individual sections should naturally merge into one another to create a holistic effect. Use the following as a checklist to ensure that you have included all of the necessary content in your abstract.

1) Identify your purpose and motivation
So your research is about rabies in Brazilian squirrels. Why is this important? You should start your abstract by explaining why people should care about this study—why is it significant to your field and perhaps to the wider world? And what is the exact purpose of your study; what are you trying to achieve? Start by answering the following questions:
- What made you decide to do this study or project?
- Why is this study important to your field or to the lay reader?
- Why should someone read your entire article?
In summary, the first section of your abstract should include the importance of the research and its impact on related research fields or on the wider scientific domain.
2) Explain the research problem you are addressing
Stating the research problem that your study addresses is the corollary to why your specific study is important and necessary. For instance, even if the issue of “rabies in Brazilian squirrels” is important, what is the problem—the “missing piece of the puzzle”—that your study helps resolve?
You can combine the problem with the motivation section, but from a perspective of organization and clarity, it is best to separate the two. Here are some precise questions to address:
- What is your research trying to better understand or what problem is it trying to solve?
- What is the scope of your study—does it try to explain something general or specific?
- What is your central claim or argument?
3) Discuss your research approach
Your specific study approach is detailed in the Methods and Materials section . You have already established the importance of the research, your motivation for studying this issue, and the specific problem your paper addresses. Now you need to discuss how you solved or made progress on this problem—how you conducted your research. If your study includes your own work or that of your team, describe that here. If in your paper you reviewed the work of others, explain this here. Did you use analytic models? A simulation? A double-blind study? A case study? You are basically showing the reader the internal engine of your research machine and how it functioned in the study. Be sure to:
- Detail your research—include methods/type of the study, your variables, and the extent of the work
- Briefly present evidence to support your claim
- Highlight your most important sources
4) Briefly summarize your results
Here you will give an overview of the outcome of your study. Avoid using too many vague qualitative terms (e.g, “very,” “small,” or “tremendous”) and try to use at least some quantitative terms (i.e., percentages, figures, numbers). Save your qualitative language for the conclusion statement. Answer questions like these:
- What did your study yield in concrete terms (e.g., trends, figures, correlation between phenomena)?
- How did your results compare to your hypothesis? Was the study successful?
- Where there any highly unexpected outcomes or were they all largely predicted?
5) State your conclusion
In the last section of your abstract, you will give a statement about the implications and limitations of the study . Be sure to connect this statement closely to your results and not the area of study in general. Are the results of this study going to shake up the scientific world? Will they impact how people see “Brazilian squirrels”? Or are the implications minor? Try not to boast about your study or present its impact as too far-reaching, as researchers and journals will tend to be skeptical of bold claims in scientific papers. Answer one of these questions:
- What are the exact effects of these results on my field? On the wider world?
- What other kind of study would yield further solutions to problems?
- What other information is needed to expand knowledge in this area?
After Completing the First Draft of Your Abstract
Revise your abstract.
The abstract, like any piece of academic writing, should be revised before being considered complete. Check it for grammatical and spelling errors and make sure it is formatted properly.
Get feedback from a peer
Getting a fresh set of eyes to review your abstract is a great way to find out whether you’ve summarized your research well. Find a reader who understands research papers but is not an expert in this field or is not affiliated with your study. Ask your reader to summarize what your study is about (including all key points of each section). This should tell you if you have communicated your key points clearly.
In addition to research peers, consider consulting with a professor or even a specialist or generalist writing center consultant about your abstract. Use any resource that helps you see your work from another perspective.
Consider getting professional editing and proofreading
While peer feedback is quite important to ensure the effectiveness of your abstract content, it may be a good idea to find an academic editor to fix mistakes in grammar, spelling, mechanics, style, or formatting. The presence of basic errors in the abstract may not affect your content, but it might dissuade someone from reading your entire study. Wordvice provides English editing services that both correct objective errors and enhance the readability and impact of your work.
Additional Abstract Rules and Guidelines
Write your abstract after completing your paper.
Although the abstract goes at the beginning of your manuscript, it does not merely introduce your research topic (that is the job of the title), but rather summarizes your entire paper. Writing the abstract last will ensure that it is complete and consistent with the findings and statements in your paper.
Keep your content in the correct order
Both questions and answers should be organized in a standard and familiar way to make the content easier for readers to absorb. Ideally, it should mimic the overall format of your essay and the classic “introduction,” “body,” and “conclusion” form, even if the parts are not neatly divided as such.
Write the abstract from scratch
Because the abstract is a self-contained piece of writing viewed separately from the body of the paper, you should write it separately as well. Never copy and paste direct quotes from the paper and avoid paraphrasing sentences in the paper. Using new vocabulary and phrases will keep your abstract interesting and free of redundancies while conserving space.
Don’t include too many details in the abstract
Again, the density of your abstract makes it incompatible with including specific points other than possibly names or locations. You can make references to terms, but do not explain or define them in the abstract. Try to strike a balance between being specific to your study and presenting a relatively broad overview of your work.
Wordvice Resources
If you think your abstract is fine now but you need input on abstract writing or require English editing services (including paper editing ), then head over to the Wordvice academic resources page, where you will find many more articles, for example on writing the Results , Methods , and Discussion sections of your manuscript, on choosing a title for your paper , or on how to finalize your journal submission with a strong cover letter .
Have a language expert improve your writing
Run a free plagiarism check in 10 minutes, automatically generate references for free.
- Knowledge Base
- Dissertation
- How to Write an Abstract | Steps & Examples
How to Write an Abstract | Steps & Examples
Published on 1 March 2019 by Shona McCombes . Revised on 10 October 2022 by Eoghan Ryan.
An abstract is a short summary of a longer work (such as a dissertation or research paper ). The abstract concisely reports the aims and outcomes of your research, so that readers know exactly what your paper is about.
Although the structure may vary slightly depending on your discipline, your abstract should describe the purpose of your work, the methods you’ve used, and the conclusions you’ve drawn.
One common way to structure your abstract is to use the IMRaD structure. This stands for:
- Introduction
Abstracts are usually around 100–300 words, but there’s often a strict word limit, so make sure to check the relevant requirements.
In a dissertation or thesis , include the abstract on a separate page, after the title page and acknowledgements but before the table of contents .
Instantly correct all language mistakes in your text
Be assured that you'll submit flawless writing. Upload your document to correct all your mistakes.
Table of contents
Abstract example, when to write an abstract, step 1: introduction, step 2: methods, step 3: results, step 4: discussion, tips for writing an abstract, frequently asked questions about abstracts.
Hover over the different parts of the abstract to see how it is constructed.
This paper examines the role of silent movies as a mode of shared experience in the UK during the early twentieth century. At this time, high immigration rates resulted in a significant percentage of non-English-speaking citizens. These immigrants faced numerous economic and social obstacles, including exclusion from public entertainment and modes of discourse (newspapers, theater, radio).
Incorporating evidence from reviews, personal correspondence, and diaries, this study demonstrates that silent films were an affordable and inclusive source of entertainment. It argues for the accessible economic and representational nature of early cinema. These concerns are particularly evident in the low price of admission and in the democratic nature of the actors’ exaggerated gestures, which allowed the plots and action to be easily grasped by a diverse audience despite language barriers.
Keywords: silent movies, immigration, public discourse, entertainment, early cinema, language barriers.
The only proofreading tool specialized in correcting academic writing
The academic proofreading tool has been trained on 1000s of academic texts and by native English editors. Making it the most accurate and reliable proofreading tool for students.
Correct my document today
You will almost always have to include an abstract when:
- Completing a thesis or dissertation
- Submitting a research paper to an academic journal
- Writing a book proposal
- Applying for research grants
It’s easiest to write your abstract last, because it’s a summary of the work you’ve already done. Your abstract should:
- Be a self-contained text, not an excerpt from your paper
- Be fully understandable on its own
- Reflect the structure of your larger work
Start by clearly defining the purpose of your research. What practical or theoretical problem does the research respond to, or what research question did you aim to answer?
You can include some brief context on the social or academic relevance of your topic, but don’t go into detailed background information. If your abstract uses specialised terms that would be unfamiliar to the average academic reader or that have various different meanings, give a concise definition.
After identifying the problem, state the objective of your research. Use verbs like “investigate,” “test,” “analyse,” or “evaluate” to describe exactly what you set out to do.
This part of the abstract can be written in the present or past simple tense but should never refer to the future, as the research is already complete.
- This study will investigate the relationship between coffee consumption and productivity.
- This study investigates the relationship between coffee consumption and productivity.
Next, indicate the research methods that you used to answer your question. This part should be a straightforward description of what you did in one or two sentences. It is usually written in the past simple tense, as it refers to completed actions.
- Structured interviews will be conducted with 25 participants.
- Structured interviews were conducted with 25 participants.
Don’t evaluate validity or obstacles here — the goal is not to give an account of the methodology’s strengths and weaknesses, but to give the reader a quick insight into the overall approach and procedures you used.
Prevent plagiarism, run a free check.
Next, summarise the main research results . This part of the abstract can be in the present or past simple tense.
- Our analysis has shown a strong correlation between coffee consumption and productivity.
- Our analysis shows a strong correlation between coffee consumption and productivity.
- Our analysis showed a strong correlation between coffee consumption and productivity.
Depending on how long and complex your research is, you may not be able to include all results here. Try to highlight only the most important findings that will allow the reader to understand your conclusions.
Finally, you should discuss the main conclusions of your research : what is your answer to the problem or question? The reader should finish with a clear understanding of the central point that your research has proved or argued. Conclusions are usually written in the present simple tense.
- We concluded that coffee consumption increases productivity.
- We conclude that coffee consumption increases productivity.
If there are important limitations to your research (for example, related to your sample size or methods), you should mention them briefly in the abstract. This allows the reader to accurately assess the credibility and generalisability of your research.
If your aim was to solve a practical problem, your discussion might include recommendations for implementation. If relevant, you can briefly make suggestions for further research.
If your paper will be published, you might have to add a list of keywords at the end of the abstract. These keywords should reference the most important elements of the research to help potential readers find your paper during their own literature searches.
Be aware that some publication manuals, such as APA Style , have specific formatting requirements for these keywords.
It can be a real challenge to condense your whole work into just a couple of hundred words, but the abstract will be the first (and sometimes only) part that people read, so it’s important to get it right. These strategies can help you get started.
Read other abstracts
The best way to learn the conventions of writing an abstract in your discipline is to read other people’s. You probably already read lots of journal article abstracts while conducting your literature review —try using them as a framework for structure and style.
You can also find lots of dissertation abstract examples in thesis and dissertation databases .

Reverse outline
Not all abstracts will contain precisely the same elements. For longer works, you can write your abstract through a process of reverse outlining.
For each chapter or section, list keywords and draft one to two sentences that summarise the central point or argument. This will give you a framework of your abstract’s structure. Next, revise the sentences to make connections and show how the argument develops.
Write clearly and concisely
A good abstract is short but impactful, so make sure every word counts. Each sentence should clearly communicate one main point.
To keep your abstract or summary short and clear:
- Avoid passive sentences: Passive constructions are often unnecessarily long. You can easily make them shorter and clearer by using the active voice.
- Avoid long sentences: Substitute longer expressions for concise expressions or single words (e.g., “In order to” for “To”).
- Avoid obscure jargon: The abstract should be understandable to readers who are not familiar with your topic.
- Avoid repetition and filler words: Replace nouns with pronouns when possible and eliminate unnecessary words.
- Avoid detailed descriptions: An abstract is not expected to provide detailed definitions, background information, or discussions of other scholars’ work. Instead, include this information in the body of your thesis or paper.
If you’re struggling to edit down to the required length, you can get help from expert editors with Scribbr’s professional proofreading services .
Check your formatting
If you are writing a thesis or dissertation or submitting to a journal, there are often specific formatting requirements for the abstract—make sure to check the guidelines and format your work correctly. For APA research papers you can follow the APA abstract format .
Checklist: Abstract
The word count is within the required length, or a maximum of one page.
The abstract appears after the title page and acknowledgements and before the table of contents .
I have clearly stated my research problem and objectives.
I have briefly described my methodology .
I have summarized the most important results .
I have stated my main conclusions .
I have mentioned any important limitations and recommendations.
The abstract can be understood by someone without prior knowledge of the topic.
You've written a great abstract! Use the other checklists to continue improving your thesis or dissertation.
An abstract is a concise summary of an academic text (such as a journal article or dissertation ). It serves two main purposes:
- To help potential readers determine the relevance of your paper for their own research.
- To communicate your key findings to those who don’t have time to read the whole paper.
Abstracts are often indexed along with keywords on academic databases, so they make your work more easily findable. Since the abstract is the first thing any reader sees, it’s important that it clearly and accurately summarises the contents of your paper.
An abstract for a thesis or dissertation is usually around 150–300 words. There’s often a strict word limit, so make sure to check your university’s requirements.
The abstract is the very last thing you write. You should only write it after your research is complete, so that you can accurately summarize the entirety of your thesis or paper.
Avoid citing sources in your abstract . There are two reasons for this:
- The abstract should focus on your original research, not on the work of others.
- The abstract should be self-contained and fully understandable without reference to other sources.
There are some circumstances where you might need to mention other sources in an abstract: for example, if your research responds directly to another study or focuses on the work of a single theorist. In general, though, don’t include citations unless absolutely necessary.
The abstract appears on its own page, after the title page and acknowledgements but before the table of contents .
Cite this Scribbr article
If you want to cite this source, you can copy and paste the citation or click the ‘Cite this Scribbr article’ button to automatically add the citation to our free Reference Generator.
McCombes, S. (2022, October 10). How to Write an Abstract | Steps & Examples. Scribbr. Retrieved 24 June 2024, from https://www.scribbr.co.uk/thesis-dissertation/abstract/
Is this article helpful?
Shona McCombes
Other students also liked, how to write a thesis or dissertation introduction, thesis & dissertation acknowledgements | tips & examples, dissertation title page.
When you choose to publish with PLOS, your research makes an impact. Make your work accessible to all, without restrictions, and accelerate scientific discovery with options like preprints and published peer review that make your work more Open.
- PLOS Biology
- PLOS Climate
- PLOS Complex Systems
- PLOS Computational Biology
- PLOS Digital Health
- PLOS Genetics
- PLOS Global Public Health
- PLOS Medicine
- PLOS Mental Health
- PLOS Neglected Tropical Diseases
- PLOS Pathogens
- PLOS Sustainability and Transformation
- PLOS Collections
- How to Write an Abstract

Expedite peer review, increase search-ability, and set the tone for your study
The abstract is your chance to let your readers know what they can expect from your article. Learn how to write a clear, and concise abstract that will keep your audience reading.
How your abstract impacts editorial evaluation and future readership
After the title , the abstract is the second-most-read part of your article. A good abstract can help to expedite peer review and, if your article is accepted for publication, it’s an important tool for readers to find and evaluate your work. Editors use your abstract when they first assess your article. Prospective reviewers see it when they decide whether to accept an invitation to review. Once published, the abstract gets indexed in PubMed and Google Scholar , as well as library systems and other popular databases. Like the title, your abstract influences keyword search results. Readers will use it to decide whether to read the rest of your article. Other researchers will use it to evaluate your work for inclusion in systematic reviews and meta-analysis. It should be a concise standalone piece that accurately represents your research.
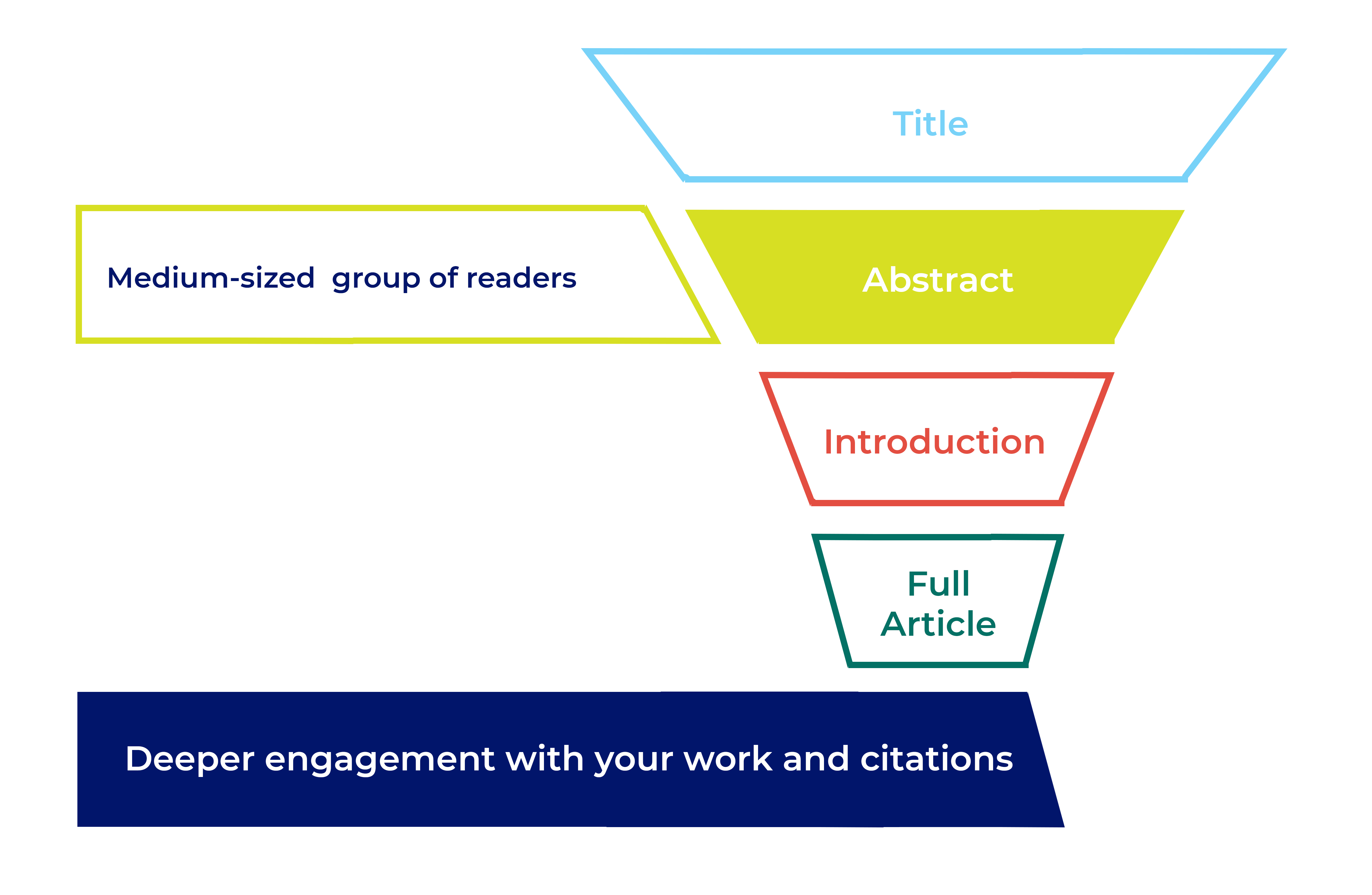
What to include in an abstract
The main challenge you’ll face when writing your abstract is keeping it concise AND fitting in all the information you need. Depending on your subject area the journal may require a structured abstract following specific headings. A structured abstract helps your readers understand your study more easily. If your journal doesn’t require a structured abstract it’s still a good idea to follow a similar format, just present the abstract as one paragraph without headings.
Background or Introduction – What is currently known? Start with a brief, 2 or 3 sentence, introduction to the research area.
Objectives or Aims – What is the study and why did you do it? Clearly state the research question you’re trying to answer.
Methods – What did you do? Explain what you did and how you did it. Include important information about your methods, but avoid the low-level specifics. Some disciplines have specific requirements for abstract methods.
- CONSORT for randomized trials.
- STROBE for observational studies
- PRISMA for systematic reviews and meta-analyses
Results – What did you find? Briefly give the key findings of your study. Include key numeric data (including confidence intervals or p values), where possible.
Conclusions – What did you conclude? Tell the reader why your findings matter, and what this could mean for the ‘bigger picture’ of this area of research.
Writing tips
The main challenge you may find when writing your abstract is keeping it concise AND convering all the information you need to.

- Keep it concise and to the point. Most journals have a maximum word count, so check guidelines before you write the abstract to save time editing it later.
- Write for your audience. Are they specialists in your specific field? Are they cross-disciplinary? Are they non-specialists? If you’re writing for a general audience, or your research could be of interest to the public keep your language as straightforward as possible. If you’re writing in English, do remember that not all of your readers will necessarily be native English speakers.
- Focus on key results, conclusions and take home messages.
- Write your paper first, then create the abstract as a summary.
- Check the journal requirements before you write your abstract, eg. required subheadings.
- Include keywords or phrases to help readers search for your work in indexing databases like PubMed or Google Scholar.
- Double and triple check your abstract for spelling and grammar errors. These kind of errors can give potential reviewers the impression that your research isn’t sound, and can make it easier to find reviewers who accept the invitation to review your manuscript. Your abstract should be a taste of what is to come in the rest of your article.

Don’t
- Sensationalize your research.
- Speculate about where this research might lead in the future.
- Use abbreviations or acronyms (unless absolutely necessary or unless they’re widely known, eg. DNA).
- Repeat yourself unnecessarily, eg. “Methods: We used X technique. Results: Using X technique, we found…”
- Contradict anything in the rest of your manuscript.
- Include content that isn’t also covered in the main manuscript.
- Include citations or references.
Tip: How to edit your work
Editing is challenging, especially if you are acting as both a writer and an editor. Read our guidelines for advice on how to refine your work, including useful tips for setting your intentions, re-review, and consultation with colleagues.
- How to Write a Great Title
- How to Write Your Methods
- How to Report Statistics
- How to Write Discussions and Conclusions
- How to Edit Your Work
The contents of the Peer Review Center are also available as a live, interactive training session, complete with slides, talking points, and activities. …
The contents of the Writing Center are also available as a live, interactive training session, complete with slides, talking points, and activities. …
There’s a lot to consider when deciding where to submit your work. Learn how to choose a journal that will help your study reach its audience, while reflecting your values as a researcher…
Have a language expert improve your writing
Run a free plagiarism check in 10 minutes, generate accurate citations for free.
- Knowledge Base
- Starting the research process
- How to Write a Research Proposal | Examples & Templates
How to Write a Research Proposal | Examples & Templates
Published on October 12, 2022 by Shona McCombes and Tegan George. Revised on November 21, 2023.
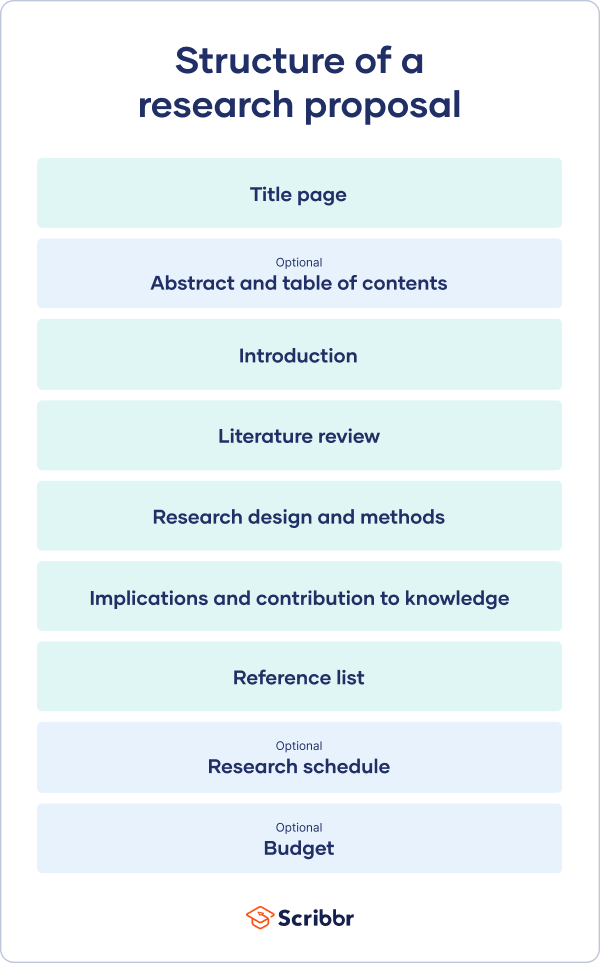
A research proposal describes what you will investigate, why it’s important, and how you will conduct your research.
The format of a research proposal varies between fields, but most proposals will contain at least these elements:
Introduction
Literature review.
- Research design
Reference list
While the sections may vary, the overall objective is always the same. A research proposal serves as a blueprint and guide for your research plan, helping you get organized and feel confident in the path forward you choose to take.
Table of contents
Research proposal purpose, research proposal examples, research design and methods, contribution to knowledge, research schedule, other interesting articles, frequently asked questions about research proposals.
Academics often have to write research proposals to get funding for their projects. As a student, you might have to write a research proposal as part of a grad school application , or prior to starting your thesis or dissertation .
In addition to helping you figure out what your research can look like, a proposal can also serve to demonstrate why your project is worth pursuing to a funder, educational institution, or supervisor.
| Show your reader why your project is interesting, original, and important. | |
| Demonstrate your comfort and familiarity with your field. Show that you understand the current state of research on your topic. | |
| Make a case for your . Demonstrate that you have carefully thought about the data, tools, and procedures necessary to conduct your research. | |
| Confirm that your project is feasible within the timeline of your program or funding deadline. |
Research proposal length
The length of a research proposal can vary quite a bit. A bachelor’s or master’s thesis proposal can be just a few pages, while proposals for PhD dissertations or research funding are usually much longer and more detailed. Your supervisor can help you determine the best length for your work.
One trick to get started is to think of your proposal’s structure as a shorter version of your thesis or dissertation , only without the results , conclusion and discussion sections.
Download our research proposal template
Prevent plagiarism. Run a free check.
Writing a research proposal can be quite challenging, but a good starting point could be to look at some examples. We’ve included a few for you below.
- Example research proposal #1: “A Conceptual Framework for Scheduling Constraint Management”
- Example research proposal #2: “Medical Students as Mediators of Change in Tobacco Use”
Like your dissertation or thesis, the proposal will usually have a title page that includes:
- The proposed title of your project
- Your supervisor’s name
- Your institution and department
The first part of your proposal is the initial pitch for your project. Make sure it succinctly explains what you want to do and why.
Your introduction should:
- Introduce your topic
- Give necessary background and context
- Outline your problem statement and research questions
To guide your introduction , include information about:
- Who could have an interest in the topic (e.g., scientists, policymakers)
- How much is already known about the topic
- What is missing from this current knowledge
- What new insights your research will contribute
- Why you believe this research is worth doing
Here's why students love Scribbr's proofreading services
Discover proofreading & editing
As you get started, it’s important to demonstrate that you’re familiar with the most important research on your topic. A strong literature review shows your reader that your project has a solid foundation in existing knowledge or theory. It also shows that you’re not simply repeating what other people have already done or said, but rather using existing research as a jumping-off point for your own.
In this section, share exactly how your project will contribute to ongoing conversations in the field by:
- Comparing and contrasting the main theories, methods, and debates
- Examining the strengths and weaknesses of different approaches
- Explaining how will you build on, challenge, or synthesize prior scholarship
Following the literature review, restate your main objectives . This brings the focus back to your own project. Next, your research design or methodology section will describe your overall approach, and the practical steps you will take to answer your research questions.
| ? or ? , , or research design? | |
| , )? ? | |
| , , , )? | |
| ? |
To finish your proposal on a strong note, explore the potential implications of your research for your field. Emphasize again what you aim to contribute and why it matters.
For example, your results might have implications for:
- Improving best practices
- Informing policymaking decisions
- Strengthening a theory or model
- Challenging popular or scientific beliefs
- Creating a basis for future research
Last but not least, your research proposal must include correct citations for every source you have used, compiled in a reference list . To create citations quickly and easily, you can use our free APA citation generator .
Some institutions or funders require a detailed timeline of the project, asking you to forecast what you will do at each stage and how long it may take. While not always required, be sure to check the requirements of your project.
Here’s an example schedule to help you get started. You can also download a template at the button below.
Download our research schedule template
| Research phase | Objectives | Deadline |
|---|---|---|
| 1. Background research and literature review | 20th January | |
| 2. Research design planning | and data analysis methods | 13th February |
| 3. Data collection and preparation | with selected participants and code interviews | 24th March |
| 4. Data analysis | of interview transcripts | 22nd April |
| 5. Writing | 17th June | |
| 6. Revision | final work | 28th July |
If you are applying for research funding, chances are you will have to include a detailed budget. This shows your estimates of how much each part of your project will cost.
Make sure to check what type of costs the funding body will agree to cover. For each item, include:
- Cost : exactly how much money do you need?
- Justification : why is this cost necessary to complete the research?
- Source : how did you calculate the amount?
To determine your budget, think about:
- Travel costs : do you need to go somewhere to collect your data? How will you get there, and how much time will you need? What will you do there (e.g., interviews, archival research)?
- Materials : do you need access to any tools or technologies?
- Help : do you need to hire any research assistants for the project? What will they do, and how much will you pay them?
If you want to know more about the research process , methodology , research bias , or statistics , make sure to check out some of our other articles with explanations and examples.
Methodology
- Sampling methods
- Simple random sampling
- Stratified sampling
- Cluster sampling
- Likert scales
- Reproducibility
Statistics
- Null hypothesis
- Statistical power
- Probability distribution
- Effect size
- Poisson distribution
Research bias
- Optimism bias
- Cognitive bias
- Implicit bias
- Hawthorne effect
- Anchoring bias
- Explicit bias
Once you’ve decided on your research objectives , you need to explain them in your paper, at the end of your problem statement .
Keep your research objectives clear and concise, and use appropriate verbs to accurately convey the work that you will carry out for each one.
I will compare …
A research aim is a broad statement indicating the general purpose of your research project. It should appear in your introduction at the end of your problem statement , before your research objectives.
Research objectives are more specific than your research aim. They indicate the specific ways you’ll address the overarching aim.
A PhD, which is short for philosophiae doctor (doctor of philosophy in Latin), is the highest university degree that can be obtained. In a PhD, students spend 3–5 years writing a dissertation , which aims to make a significant, original contribution to current knowledge.
A PhD is intended to prepare students for a career as a researcher, whether that be in academia, the public sector, or the private sector.
A master’s is a 1- or 2-year graduate degree that can prepare you for a variety of careers.
All master’s involve graduate-level coursework. Some are research-intensive and intend to prepare students for further study in a PhD; these usually require their students to write a master’s thesis . Others focus on professional training for a specific career.
Critical thinking refers to the ability to evaluate information and to be aware of biases or assumptions, including your own.
Like information literacy , it involves evaluating arguments, identifying and solving problems in an objective and systematic way, and clearly communicating your ideas.
The best way to remember the difference between a research plan and a research proposal is that they have fundamentally different audiences. A research plan helps you, the researcher, organize your thoughts. On the other hand, a dissertation proposal or research proposal aims to convince others (e.g., a supervisor, a funding body, or a dissertation committee) that your research topic is relevant and worthy of being conducted.
Cite this Scribbr article
If you want to cite this source, you can copy and paste the citation or click the “Cite this Scribbr article” button to automatically add the citation to our free Citation Generator.
McCombes, S. & George, T. (2023, November 21). How to Write a Research Proposal | Examples & Templates. Scribbr. Retrieved June 27, 2024, from https://www.scribbr.com/research-process/research-proposal/
Is this article helpful?
Shona McCombes
Other students also liked, how to write a problem statement | guide & examples, writing strong research questions | criteria & examples, how to write a literature review | guide, examples, & templates, what is your plagiarism score.

Magnum Proofreading Services
- Jake Magnum
- Jan 2, 2021
Writing an Abstract for a Research Paper: Guidelines, Examples, and Templates
There are six steps to writing a standard abstract. (1) Begin with a broad statement about your topic. Then, (2) state the problem or knowledge gap related to this topic that your study explores. After that, (3) describe what specific aspect of this problem you investigated, and (4) briefly explain how you went about doing this. After that, (5) describe the most meaningful outcome(s) of your study. Finally, (6) close your abstract by explaining the broad implication(s) of your findings.
In this article, I present step-by-step guidelines for writing an abstract for an academic paper. These guidelines are fo llowed by an example of a full abstract that follows these guidelines and a few fill-in-the-blank templates that you can use to write your own abstract.
Guidelines for Writing an Abstract
The basic structure of an abstract is illustrated below.

A standard abstract starts with a very general statement and becomes more specific with each sentence that follows until once again making a broad statement about the study’s implications at the end. Altogether, a standard abstract has six functions, which are described in detail below.
Start by making a broad statement about your topic.
The first sentence of your abstract should briefly describe a problem that is of interest to your readers. When writing this first sentence, you should think about who comprises your target audience and use terms that will appeal to this audience. If your opening sentence is too broad, it might lose the attention of potential readers because they will not know if your study is relevant to them.
Too broad : Maintaining an ideal workplace environment has a positive effect on employees.
The sentence above is so broad that it will not grab the reader’s attention. While it gives the reader some idea of the area of study, it doesn’t provide any details about the author’s topic within their research area. This can be fixed by inserting some keywords related to the topic (these are underlined in the revised example below).
Improved : Keeping the workplace environment at an ideal temperature positively affects the overall health of employees.
The revised sentence is much better, as it expresses two points about the research topic—namely, (i) what aspect of workplace environment was studied, (ii) what aspect of employees was observed. The mention of these aspects of the research will draw the attention of readers who are interested in them.
Describe the general problem that your paper addresses.
After describing your topic in the first sentence, you can then explain what aspect of this topic has motivated your research. Often, authors use this part of the abstract to describe the research gap that they identified and aimed to fill. These types of sentences are often characterized by the use of words such as “however,” “although,” “despite,” and so on.
However, a comprehensive understanding of how different workplace bullying experiences are associated with absenteeism is currently lacking.
The above example is typical of a sentence describing the problem that a study intends to tackle. The author has noticed that there is a gap in the research, and they briefly explain this gap here.
Although it has been established that quantity and quality of sleep can affect different types of task performance and personal health, the interactions between sleep habits and workplace behaviors have received very little attention.
The example above illustrates a case in which the author has accomplished two tasks with one sentence. The first part of the sentence (up until the comma) mentions the general topic that the research fits into, while the second part (after the comma) describes the general problem that the research addresses.
Express the specific problem investigated in your paper.
After describing the general problem that motivated your research, the next sentence should express the specific aspect of the problem that you investigated. Sentences of this type are often indicated by the use of phrases like “the purpose of this research is to,” “this paper is intended to,” or “this work aims to.”
Uninformative : However, a comprehensive understanding of how different workplace bullying experiences are associated with absenteeism is currently lacking. The present article aimed to provide new insights into the relationship between workplace bullying and absenteeism .
The second sentence in the above example is a mere rewording of the first sentence. As such, it adds nothing to the abstract. The second sentence should be more specific than the preceding one.
Improved : However, a comprehensive understanding of how different workplace bullying experiences are associated with absenteeism is currently lacking. The present article aimed to define various subtypes of workplace bullying and determine which subtypes tend to lead to absenteeism .
The second sentence of this passage is much more informative than in the previous example. This sentence lets the reader know exactly what they can expect from the full research article.
Explain how you attempted to resolve your study’s specific problem.
In this part of your abstract, you should attempt to describe your study’s methodology in one or two sentences. As such, you must be sure to include only the most important information about your method. At the same time, you must also be careful not to be too vague.
Too vague : We conducted multiple tests to examine changes in various factors related to well-being.
This description of the methodology is too vague. Instead of merely mentioning “tests” and “factors,” the author should note which specific tests were run and which factors were assessed.
Improved : Using data from BHIP completers, we conducted multiple one-way multivariate analyses of variance and follow-up univariate t-tests to examine changes in physical and mental health, stress, energy levels, social satisfaction, self-efficacy, and quality of life.
This sentence is very well-written. It packs a lot of specific information about the method into a single sentence. Also, it does not describe more details than are needed for an abstract.
Briefly tell the reader what you found by carrying out your study.
This is the most important part of the abstract—the other sentences in the abstract are there to explain why this one is relevant. When writing this sentence, imagine that someone has asked you, “What did you find in your research?” and that you need to answer them in one or two sentences.
Too vague : Consistently poor sleepers had more health risks and medical conditions than consistently optimal sleepers.
This sentence is okay, but it would be helpful to let the reader know which health risks and medical conditions were related to poor sleeping habits.
Improved : Consistently poor sleepers were more likely than consistently optimal sleepers to suffer from chronic abdominal pain, and they were at a higher risk for diabetes and heart disease.
This sentence is better, as the specific health conditions are named.
Finally, describe the major implication(s) of your study.
Most abstracts end with a short sentence that explains the main takeaway(s) that you want your audience to gain from reading your paper. Often, this sentence is addressed to people in power (e.g., employers, policymakers), and it recommends a course of action that such people should take based on the results.
Too broad : Employers may wish to make use of strategies that increase employee health.
This sentence is too broad to be useful. It does not give employers a starting point to implement a change.
Improved : Employers may wish to incorporate sleep education initiatives as part of their overall health and wellness strategies.
This sentence is better than the original, as it provides employers with a starting point—specifically, it invites employers to look up information on sleep education programs.
Abstract Example
The abstract produced here is from a paper published in Electronic Commerce Research and Applications . I have made slight alterations to the abstract so that this example fits the guidelines given in this article.
(1) Gamification can strengthen enjoyment and productivity in the workplace. (2) Despite this, research on gamification in the work context is still limited. (3) In this study, we investigated the effect of gamification on the workplace enjoyment and productivity of employees by comparing employees with leadership responsibilities to those without leadership responsibilities. (4) Work-related tasks were gamified using the habit-tracking game Habitica, and data from 114 employees were gathered using an online survey. (5) The results illustrated that employees without leadership responsibilities used work gamification as a trigger for self-motivation, whereas employees with leadership responsibilities used it to improve their health. (6) Work gamification positively affected work enjoyment for both types of employees and positively affected productivity for employees with leadership responsibilities. (7) Our results underline the importance of taking work-related variables into account when researching work gamification.
In Sentence (1), the author makes a broad statement about their topic. Notice how the nouns used (“gamification,” “enjoyment,” “productivity”) are quite general while still indicating the focus of the paper. The author uses Sentence (2) to very briefly state the problem that the research will address.
In Sentence (3), the author explains what specific aspects of the problem mentioned in Sentence (2) will be explored in the present work. Notice that the mention of leadership responsibilities makes Sentence (3) more specific than Sentence (2). Sentence (4) gets even more specific, naming the specific tools used to gather data and the number of participants.
Sentences (5) and (6) are similar, with each sentence describing one of the study’s main findings. Then, suddenly, the scope of the abstract becomes quite broad again in Sentence (7), which mentions “work-related variables” instead of a specific variable and “researching” instead of a specific kind of research.
Abstract Templates
Copy and paste any of the paragraphs below into a word processor. Then insert the appropriate information to produce an abstract for your research paper.
Template #1
Researchers have established that [Make a broad statement about your area of research.] . However, [Describe the knowledge gap that your paper addresses.] . The goal of this paper is to [Describe the purpose of your paper.] . The achieve this goal, we [Briefly explain your methodology.] . We found that [Indicate the main finding(s) of your study; you may need two sentences to do this.] . [Provide a broad implication of your results.] .
Template #2
It is well-understood that [Make a broad statement about your area of research.] . Despite this, [Describe the knowledge gap that your paper addresses.] . The current research aims to [Describe the purpose of your paper.] . To accomplish this, we [Briefly explain your methodology.] . It was discovered that [Indicate the main finding(s) of your study; you may need two sentences to do this.] . [Provide a broad implication of your results.] .
Template #3
Extensive research indicates that [Make a broad statement about your area of research.] . Nevertheless, [Describe the knowledge gap that your paper addresses.] . The present work is intended to [Describe the purpose of your paper.] . To this end, we [Briefly explain your methodology.] . The results revealed that [Indicate the main finding(s) of your study; you may need two sentences to do this.] . [Provide a broad implication of your results.] .
- How to Write an Abstract
Related Posts
How to Write a Research Paper in English: A Guide for Non-native Speakers
How to Write an Abstract Quickly
Using the Present Tense and Past Tense When Writing an Abstract
- USC Libraries
- Research Guides
Organizing Your Social Sciences Research Paper
- 3. The Abstract
- Purpose of Guide
- Design Flaws to Avoid
- Independent and Dependent Variables
- Glossary of Research Terms
- Reading Research Effectively
- Narrowing a Topic Idea
- Broadening a Topic Idea
- Extending the Timeliness of a Topic Idea
- Academic Writing Style
- Applying Critical Thinking
- Choosing a Title
- Making an Outline
- Paragraph Development
- Research Process Video Series
- Executive Summary
- The C.A.R.S. Model
- Background Information
- The Research Problem/Question
- Theoretical Framework
- Citation Tracking
- Content Alert Services
- Evaluating Sources
- Primary Sources
- Secondary Sources
- Tiertiary Sources
- Scholarly vs. Popular Publications
- Qualitative Methods
- Quantitative Methods
- Insiderness
- Using Non-Textual Elements
- Limitations of the Study
- Common Grammar Mistakes
- Writing Concisely
- Avoiding Plagiarism
- Footnotes or Endnotes?
- Further Readings
- Generative AI and Writing
- USC Libraries Tutorials and Other Guides
- Bibliography
An abstract summarizes, usually in one paragraph of 300 words or less, the major aspects of the entire paper in a prescribed sequence that includes: 1) the overall purpose of the study and the research problem(s) you investigated; 2) the basic design of the study; 3) major findings or trends found as a result of your analysis; and, 4) a brief summary of your interpretations and conclusions.
Writing an Abstract. The Writing Center. Clarion University, 2009; Writing an Abstract for Your Research Paper. The Writing Center, University of Wisconsin, Madison; Koltay, Tibor. Abstracts and Abstracting: A Genre and Set of Skills for the Twenty-first Century . Oxford, UK: Chandos Publishing, 2010;
Importance of a Good Abstract
Sometimes your professor will ask you to include an abstract, or general summary of your work, with your research paper. The abstract allows you to elaborate upon each major aspect of the paper and helps readers decide whether they want to read the rest of the paper. Therefore, enough key information [e.g., summary results, observations, trends, etc.] must be included to make the abstract useful to someone who may want to examine your work.
How do you know when you have enough information in your abstract? A simple rule-of-thumb is to imagine that you are another researcher doing a similar study. Then ask yourself: if your abstract was the only part of the paper you could access, would you be happy with the amount of information presented there? Does it tell the whole story about your study? If the answer is "no" then the abstract likely needs to be revised.
Farkas, David K. “A Scheme for Understanding and Writing Summaries.” Technical Communication 67 (August 2020): 45-60; How to Write a Research Abstract. Office of Undergraduate Research. University of Kentucky; Staiger, David L. “What Today’s Students Need to Know about Writing Abstracts.” International Journal of Business Communication January 3 (1966): 29-33; Swales, John M. and Christine B. Feak. Abstracts and the Writing of Abstracts . Ann Arbor, MI: University of Michigan Press, 2009.
Structure and Writing Style
I. Types of Abstracts
To begin, you need to determine which type of abstract you should include with your paper. There are four general types.
Critical Abstract A critical abstract provides, in addition to describing main findings and information, a judgment or comment about the study’s validity, reliability, or completeness. The researcher evaluates the paper and often compares it with other works on the same subject. Critical abstracts are generally 400-500 words in length due to the additional interpretive commentary. These types of abstracts are used infrequently.
Descriptive Abstract A descriptive abstract indicates the type of information found in the work. It makes no judgments about the work, nor does it provide results or conclusions of the research. It does incorporate key words found in the text and may include the purpose, methods, and scope of the research. Essentially, the descriptive abstract only describes the work being summarized. Some researchers consider it an outline of the work, rather than a summary. Descriptive abstracts are usually very short, 100 words or less. Informative Abstract The majority of abstracts are informative. While they still do not critique or evaluate a work, they do more than describe it. A good informative abstract acts as a surrogate for the work itself. That is, the researcher presents and explains all the main arguments and the important results and evidence in the paper. An informative abstract includes the information that can be found in a descriptive abstract [purpose, methods, scope] but it also includes the results and conclusions of the research and the recommendations of the author. The length varies according to discipline, but an informative abstract is usually no more than 300 words in length.
Highlight Abstract A highlight abstract is specifically written to attract the reader’s attention to the study. No pretense is made of there being either a balanced or complete picture of the paper and, in fact, incomplete and leading remarks may be used to spark the reader’s interest. In that a highlight abstract cannot stand independent of its associated article, it is not a true abstract and, therefore, rarely used in academic writing.
II. Writing Style
Use the active voice when possible , but note that much of your abstract may require passive sentence constructions. Regardless, write your abstract using concise, but complete, sentences. Get to the point quickly and always use the past tense because you are reporting on a study that has been completed.
Abstracts should be formatted as a single paragraph in a block format and with no paragraph indentations. In most cases, the abstract page immediately follows the title page. Do not number the page. Rules set forth in writing manual vary but, in general, you should center the word "Abstract" at the top of the page with double spacing between the heading and the abstract. The final sentences of an abstract concisely summarize your study’s conclusions, implications, or applications to practice and, if appropriate, can be followed by a statement about the need for additional research revealed from the findings.
Composing Your Abstract
Although it is the first section of your paper, the abstract should be written last since it will summarize the contents of your entire paper. A good strategy to begin composing your abstract is to take whole sentences or key phrases from each section of the paper and put them in a sequence that summarizes the contents. Then revise or add connecting phrases or words to make the narrative flow clearly and smoothly. Note that statistical findings should be reported parenthetically [i.e., written in parentheses].
Before handing in your final paper, check to make sure that the information in the abstract completely agrees with what you have written in the paper. Think of the abstract as a sequential set of complete sentences describing the most crucial information using the fewest necessary words. The abstract SHOULD NOT contain:
- A catchy introductory phrase, provocative quote, or other device to grab the reader's attention,
- Lengthy background or contextual information,
- Redundant phrases, unnecessary adverbs and adjectives, and repetitive information;
- Acronyms or abbreviations,
- References to other literature [say something like, "current research shows that..." or "studies have indicated..."],
- Using ellipticals [i.e., ending with "..."] or incomplete sentences,
- Jargon or terms that may be confusing to the reader,
- Citations to other works, and
- Any sort of image, illustration, figure, or table, or references to them.
Abstract. Writing Center. University of Kansas; Abstract. The Structure, Format, Content, and Style of a Journal-Style Scientific Paper. Department of Biology. Bates College; Abstracts. The Writing Center. University of North Carolina; Borko, Harold and Seymour Chatman. "Criteria for Acceptable Abstracts: A Survey of Abstracters' Instructions." American Documentation 14 (April 1963): 149-160; Abstracts. The Writer’s Handbook. Writing Center. University of Wisconsin, Madison; Hartley, James and Lucy Betts. "Common Weaknesses in Traditional Abstracts in the Social Sciences." Journal of the American Society for Information Science and Technology 60 (October 2009): 2010-2018; Koltay, Tibor. Abstracts and Abstracting: A Genre and Set of Skills for the Twenty-first Century. Oxford, UK: Chandos Publishing, 2010; Procter, Margaret. The Abstract. University College Writing Centre. University of Toronto; Riordan, Laura. “Mastering the Art of Abstracts.” The Journal of the American Osteopathic Association 115 (January 2015 ): 41-47; Writing Report Abstracts. The Writing Lab and The OWL. Purdue University; Writing Abstracts. Writing Tutorial Services, Center for Innovative Teaching and Learning. Indiana University; Koltay, Tibor. Abstracts and Abstracting: A Genre and Set of Skills for the Twenty-First Century . Oxford, UK: 2010; Writing an Abstract for Your Research Paper. The Writing Center, University of Wisconsin, Madison.
Writing Tip
Never Cite Just the Abstract!
Citing to just a journal article's abstract does not confirm for the reader that you have conducted a thorough or reliable review of the literature. If the full-text is not available, go to the USC Libraries main page and enter the title of the article [NOT the title of the journal]. If the Libraries have a subscription to the journal, the article should appear with a link to the full-text or to the journal publisher page where you can get the article. If the article does not appear, try searching Google Scholar using the link on the USC Libraries main page. If you still can't find the article after doing this, contact a librarian or you can request it from our free i nterlibrary loan and document delivery service .
- << Previous: Research Process Video Series
- Next: Executive Summary >>
- Last Updated: Jun 18, 2024 10:45 AM
- URL: https://libguides.usc.edu/writingguide
Five Steps to a Brilliant Abstract
by Dr. Jo Koster, Winthrop University
Humanities scholars and students aren’t usually taught to write abstracts like our friends in the natural and social sciences are. That’s because in the humanities, full pieces of discourse are preferred to short, condensed summaries. But in many cases you will NEED to write an abstract for your work—and a lot of what your colleagues in other disciplines know can help you.
Let’s start with the basic questions.
What is a descriptive abstract?
A descriptive abstract is the summary of work you have already completed or work you are proposing. It is not the same thing as the introduction to your work. The abstract should give readers a short, concise snapshot of the work as a whole—not just how it starts. Remember that the readers of your abstract will sometimes not read the paper as a whole, so in this short document you need to give them an overall picture of your work. If you are writing an abstract as a proposal for your research—in other words, as a request for permission to write a paper—the abstract serves to predict the kind of paper you hope to write.
What’s different about a conference paper (or informative) abstract?
A conference abstract is one you submit to have your paper considered for presentation at a professional conference (CURAH maintains a growing list of these opportunities ). The conference organizers will specify the length — rarely be more than 500 words (just short of two double-spaced pages). In an ideal world, you write your abstract after the actual paper is completed, but in some cases you may write an abstract for a paper you haven’t yet written—especially if the conference is some time away. Because the conference review committee will usually read the abstract and not your actual paper, you need to think of it as an independent document, aimed at that specific committee and connecting solidly with the theme of the conference. You may want to pick up phrasing from the conference title or call for papers in the abstract to reinforce this connection. Examine the call for papers carefully; it will specify the length of the abstract, special formatting requirements, whether the abstract will be published in the conference bulletin or proceedings, etc. Abstracts that do not meet the specified format are usually rejected early in the proceedings, so pay attention to each conference’s rules!
How wedded are you to the abstract you submit?
An abstract is a promissory note. That is, you are promising that you can and will produce the goods in the paper. Particularly in the case of a conference abstract, the organizers will make up a session based on the contents of the abstract. If you propose a paper that says you will use Foucault to comment on post-colonialism in Heat and Dust” and then show up with a paper on “Metaphors for Spring in A Bend in the River,” your paper may not fit the session where it was slotted, and you’ll look silly—and those organizers may not ask you back. While some divergence from the promised topic is acceptable (and probably inevitable if you haven’t written the paper when you submit the abstract), you need to produce a paper that’s within shouting distance of your original topic for the sake of keeping your promise.
The Five Step Process
Descriptive abstracts are usually only 100-250 words, so they must be pared down to the essentials. Typically, a descriptive abstract answers these questions:
Why did you choose this study or project? What did/will you do and how? What did you/do you hope to find? (For a completed work) What do your findings mean?
Step 1: A catchy title
Which paper would you rather go hear at a conference? ‘Issues of Heteronormativity and Gender Performance In Twain’s Novels” or “Come Back to the Raft, Huck Honey”?
Your title should be informative and focused, indicating the problem and your general approach. It’s very fashionable in the humanities to have titles featuring a catchy phrase, a colon, and then an explanation of the title. While snappy titles may help your abstract be noticed, it’s really what comes after the colon that sells the abstract, so pay attention to it. “All the World’s a Ship: Race and Ethnicity in Moby Dick” catches the eye, but “Melville’s Deconstruction of Ethnicity in the ‘Midnight, Forecastle’ Episode of Moby Dick” tells readers much more specifically what you’re promising to deliver.
Step 2: A snappy context sentence (or sentences)
The abstract should begin with a clear sense of the research question you have framed. Often writers set this up as a problem: “Although some recent scholars claim to have identified Shakespeare’s lost play Cardenio, that attribution is still not accepted.
Step 3: Introduce your argument (don’t just copy your thesis statement).
If you began with a problem, you can pose your argument as the solution: “In this paper I use the records of the Worshipful Company of Stationers, London’s chief publishing organization, to show that the play identified by Charles Hamilton in 1990 is not actually the play Shakespeare’s company mounted in 1613.” It’s perfectly legit to use “I” in sentences referring to your argument.
Step 4: Add some sentences describing how you make your argument.
It always helps when you identify the theoretical or methodological school that you are using to approach your question or position yourself within an ongoing debate. This helps readers situate your ideas in the larger conversations of your discipline. For instance, “The debate among Folsom, McGann, and Stallybrass over the notion of database as a genre (PMLA 122.5, Fall 2007) suggests that….” or “Using the definition of dataclouds proposed by Johnson-Eilola (2005), I will argue that…”
Finally, briefly state your conclusion.
“ Through analyzing Dickinson’s use of metaphor, I demonstrate that she systematically transformed Watt’s hymnal tropes as a way of asserting her own doctrinal truths. This transformation…”
Not everyone agrees how much jargon should be included in an abstract. My best advice is to add any technical terms you need, but don’t put in jargon for jargon’s sake or just to make it look like you are an expert (this especially extends to (post)modernizing your words or other typographical excrescences).
Special for conference papers:
To the basic requirements of the descriptive abstract, a conference paper abstract should also include a few sentences about how the proposed paper fits in the theme of the conference. For instance, a call for papers for a session on “Science and Literature in the 19th Century” at a conference entitled “(Dis)Junctions” requested “critical works on the interaction between scientific writing and literature in the 19th century. How did scientific discoveries, theories and assumptions (for example, in medicine and psychology, but not limited to these) influence contemporaneous fiction?” If you were submitting a paper to this session, you would want to have a sentence or two about the theories you were discussing and name the particular works where you would identify their influence. If you can work the words “join” or “junction” (or “disjunction”) into your title or abstract, you’ll increase your chance of having the paper accepted, since you’re showing clearly how the paper fits the theme of the session.
Step 5: Show the conference organizers or editors that you’re a pro.
Tell them your essay is a finished work (even if it’s only complete in your head!). It’s also considered good in a conference abstract to conclude with a sentence about your presentation, since the great horror of session chairs is the paper that runs far too long (or embarrassingly too short). Organizers also need to know if you need any special technology to present the paper. So a a much-appreciated professional touch is concluding passage such as, “My paper is complete and can be presented in 20 minutes. I will bring bring video clips on a portable drive but will need a computer, projector, and Internet access to show all my materials.”
Be professional!
Double-check your abstract to make sure it meets the length requirements. Make sure it’s edited and documented. And above all, make sure it’s submitted on time.
Here is a video version of this page, taking you from the call for papers to the finished abstract.
Check out these other guides from CURAH:
- How to write a proposal
- How to make a poster
- A list of regional and national conferences where you can present your work
Acknowledgements:
Illustrated by Ian MacInnes Thanks to Dr. Leslie Bickford for her sample abstract
I consulted and borrowed material from the following websites in preparing these suggestions:
www.unc.edu/depts/wcweb/handouts/abstracts.html www.linguistics.ucsb.edu/faculty/bucholtz/sociocultural/abstracttips.html www.academic-conferences.org/abstract-guidelines.htm ceca.icom.museum/ dbase upl/writinganabstract.pdf ling.wisc.edu/macaulay/800.abstracts.html writingcenter.unlv.edu/writing/abstract.html www.lightbluetouchpaper.org/2007/03/14/how-not-to-write-an-abstract/ webapp.comcol.umass.edu/msc/absGuidelines.aspx www.oberlin.edu/history/Honors/prospectus.html www.english.eku.edu/ma/scholarlythesis.php
The Arts and Humanities Division of the Council on Undergraduate Research
- How to Cite
- Language & Lit
- Rhyme & Rhythm
- The Rewrite
- Search Glass
How to Write an Abstract for a Proposal
A proposal paper sets out your reasoning for the study, justifies the research and explains your intended methods. Dissertations and other graduate-level research often require proposals, or you may create one to apply for grant money. An abstract summarizes the information in the proposal. An effective abstract can make the difference between a positive or negative response to the proposal.
Write About the Introduction and Problem
A strong abstract touches on all the sections in the proposal, including the introduction, where you should give some information about the issue and why you chose it. While you do not want to go into detail about the problem, you need to state what issue your project will address, such as the high dropout rate for sophomores at a college. If you find you cannot focus your abstract on a single problem, your research may be too broad.
Summarize the Background and Focus
A proposal identifies a reason for the project, so the abstract also needs to establish how this project fulfills a need. You may indicate how your plan differs from previous research or fills a void in past research while summarizing information included in the literature review portion of your paper. Include a brief explanation of the project's objectives, the research or other material you will rely on in the paper and in your proposed thesis.
Explain the Methods and Conclusions
The abstract should include some general information about the procedures for your project. Explain if you will use qualitative, quantitative or mixed measures and why. What type of sample and procedures will you use to obtain your data? Add a sentence at the end of the abstract to indicate the conclusion you expect to draw from the project and the implications of the results, which will create a sense of closure for the document. Remember, the abstract is a summary of material in the paper, so only include information in the abstract that will also appear in the actual paper.
Follow Proper Formatting
First person point of view -- "I" and "my" -- are usually acceptable in APA proposals, but you should double check your field's style guide. After finishing a draft, revise your abstract to create concise language, keeping the abstract to a maximum of 250 words. Find examples of acceptable abstracts from your field and institution to use as models. If you write the abstract before finishing the proposal, review it once you have completed the paper to make sure the abstract summarizes the ideas you have presented. Insert a page break after the title page and place the abstract there, including the running head and page number in the header.
- University of Utah: How to Write a Graduate Proposal
- University of Nevada, Las Vegas: Writing Tips: How to Write an Abstract
- Rochester Institute of Technology: Writing a Successful Proposal
- University of Oregon: How to Write a Proposal Abstract
- Chapman University: How to Write an Abstract
- Purdue University: APA Stylistics: Basics
- Purdue University: General Format
Kristie Sweet has been writing professionally since 1982, most recently publishing for various websites on topics like health and wellness, and education. She holds a Master of Arts in English from the University of Northern Colorado.
Sample abstracts
Reprocessing used nuclear fuel (UNF) is crucial to the completion of a closed fuel cycle and would reduce the volume of waste produced during nuclear power production. Pyroprocessing is a promising reprocessing technique as it offers pure forms of product recovery. A limiting issue with pyroprocessing, however, is the inability to monitor concentrations of chemical species inside the electrorefiner. As with many nuclear processes, safe guards and monitoring become increasingly important; therefore, development of real - time monitoring techniques for various chemical species may allow for commercialization of this recycling process [1 - 5]. The focus of the proposed research is to develop accurate diffusion coefficients for Yttrium, a fission product found in UNF, in molten salt conditions through Cyclic Voltammetry (CV). Quantification of the diffusion coefficient will allow current measurements from inside the melt to be directly related to species concentration. With the diffusion coefficients, in - situ CV would then facilitate real - time monitoring of chemical concentrations.
This project aims to analyze the social and cultural effects of the Iranian Revolution through primary source material and interviews with those directly affected by the revolution. Iran’s political seclusion and its animosity toward the West has limited the voices and perspectives available to an American audience. Moreover, the attitude of the West towards Iran since the revolution has been myopic and often marred by political perspectives. The objective of this project will be to bring those voices and stories to light, putting a greater focus on the experiences of individuals who lived through the Revolution. These stories will be presented in a digital medium (film and web) in order to bring these voices and perspectives to an American audience.
Thank you for visiting nature.com. You are using a browser version with limited support for CSS. To obtain the best experience, we recommend you use a more up to date browser (or turn off compatibility mode in Internet Explorer). In the meantime, to ensure continued support, we are displaying the site without styles and JavaScript.
- View all journals
- Explore content
- About the journal
- Publish with us
- Sign up for alerts
- Published: 25 June 2024
Strain-specific gut microbial signatures in type 2 diabetes identified in a cross-cohort analysis of 8,117 metagenomes
- Zhendong Mei ORCID: orcid.org/0000-0001-6235-8647 1 , 2 na1 ,
- Fenglei Wang ORCID: orcid.org/0000-0002-3850-2482 2 , 3 na1 ,
- Amrisha Bhosle 2 , 4 ,
- Danyue Dong ORCID: orcid.org/0009-0003-3256-2876 1 , 2 ,
- Raaj Mehta 2 , 5 , 6 , 7 ,
- Andrew Ghazi 2 , 4 ,
- Yancong Zhang ORCID: orcid.org/0000-0002-2768-2975 2 , 4 ,
- Yuxi Liu ORCID: orcid.org/0000-0003-2484-151X 1 , 2 , 8 ,
- Ehud Rinott 9 ,
- Siyuan Ma 10 ,
- Eric B. Rimm ORCID: orcid.org/0000-0002-1402-7250 1 , 3 , 8 ,
- Martha Daviglus 11 ,
- Walter C. Willett 1 , 3 , 8 ,
- Rob Knight ORCID: orcid.org/0000-0002-0975-9019 12 , 13 , 14 ,
- Frank B. Hu 1 , 3 , 8 ,
- Qibin Qi ORCID: orcid.org/0000-0002-2687-1758 3 , 15 ,
- Andrew T. Chan ORCID: orcid.org/0000-0001-7284-6767 2 , 5 , 6 ,
- Robert D. Burk ORCID: orcid.org/0000-0002-8376-8458 15 , 16 , 17 , 18 ,
- Meir J. Stampfer 1 , 3 , 8 ,
- Iris Shai 3 , 19 ,
- Robert C. Kaplan 15 , 20 ,
- Curtis Huttenhower ORCID: orcid.org/0000-0002-1110-0096 2 , 4 , 21 , 22 &
- Dong D. Wang ORCID: orcid.org/0000-0002-0897-3048 1 , 2 , 3
Nature Medicine ( 2024 ) Cite this article
1741 Accesses
187 Altmetric
Metrics details
- Clinical microbiology
- Type 2 diabetes
The association of gut microbial features with type 2 diabetes (T2D) has been inconsistent due in part to the complexity of this disease and variation in study design. Even in cases in which individual microbial species have been associated with T2D, mechanisms have been unable to be attributed to these associations based on specific microbial strains. We conducted a comprehensive study of the T2D microbiome, analyzing 8,117 shotgun metagenomes from 10 cohorts of individuals with T2D, prediabetes, and normoglycemic status in the United States, Europe, Israel and China. Dysbiosis in 19 phylogenetically diverse species was associated with T2D (false discovery rate < 0.10), for example, enriched Clostridium bolteae and depleted Butyrivibrio crossotus . These microorganisms also contributed to community-level functional changes potentially underlying T2D pathogenesis, for example, perturbations in glucose metabolism. Our study identifies within-species phylogenetic diversity for strains of 27 species that explain inter-individual differences in T2D risk, such as Eubacterium rectale . In some cases, these were explained by strain-specific gene carriage, including loci involved in various mechanisms of horizontal gene transfer and novel biological processes underlying metabolic risk, for example, quorum sensing. In summary, our study provides robust cross-cohort microbial signatures in a strain-resolved manner and offers new mechanistic insights into T2D.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
24,99 € / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
195,33 € per year
only 16,28 € per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
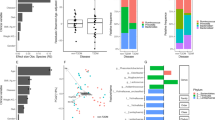

Revealing links between gut microbiome and its fungal community in Type 2 Diabetes Mellitus among Emirati subjects: A pilot study
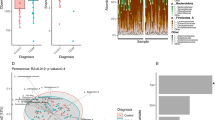

Functional alterations and predictive capacity of gut microbiome in type 2 diabetes
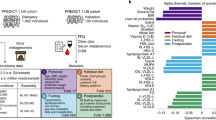
Microbiome connections with host metabolism and habitual diet from 1,098 deeply phenotyped individuals
Data availability.
The individual-level raw shotgun sequencing data and metadata have been deposited in the European Nucleotide Archive with accession codes PRJEB37249 , PRJEB38742 , PRJEB41311 and PRJEB46098 for the Fromentin_2022 dataset (MetaCardis); the Sequence Read Archive (SRA) under accession code ERP002469 for the Karlsson_2013 dataset; the NCBI SRA under accession numbers SRA045646 and SRA050230 for the Qin_2012 dataset (Shenzhen cohort); the China NGDC Genome Sequence Archive: HRA000020 or EGA: EGAS00001004480 for the Wu_2020 dataset; and the China Nucleotide Sequence Archive (CNSA) with the dataset identifier CNP0000175 for the Zhong_2019 dataset (Suzhou cohort). The shotgun metagenomic sequencing data from the Nurses’ Health Study II (NHSII) and Health Professionals Follow-up Study (HPFS) are publicly available at the BIOM-Mass Data Portal ( https://biom-mass.org/ ; project names: HPFS and MBS). Due to the gaining of informed consent from the participants, all of the individual-level phenotype data from NHSII and HPFS are available via a request for external collaboration and upon approval of a letter of intent and a research proposal. Details on how to request external collaboration with NHSII and HPFS can be found at https://nurseshealthstudy.org/researchers (contact principal investigator: A. H. Eliassen, email: [email protected]) and https://sites.sph.harvard.edu/hpfs/for-collaborators/ (contact principal investigator L. Mucci, email: [email protected]). The individual-level metadata in the Hispanic Community Health Study/Study of Latinos (HCHS/SOL) are archived at the National Institutes of Health repositories dbGap (study accession: phs000810.v2.p2 ) and BIOLINCC (accession number: HLB01141423a). Shotgun metagenomic sequencing data from the HCHS/SOL samples described in this study are deposited in QIITA (study ID: 11666). HCHS/SOL has established a process for the scientific community to apply for access to participant data and materials, with such requests reviewed by the project’s Steering Committee. These policies are described at https://sites.cscc.unc.edu/hchs/ (contact HCHS/SOL at [email protected]). The DIRECT-PLUS Study recruited participants in Israel and was designed as a clinical trial. That study used only baseline, pre-randomization data from the DIRECT-PLUS Study for an observational analysis. Due to gaining of informed consent from the participants, the individual-level de-identified metadata and metagenomic sequencing data in the DIRECT-PLUS Study will be available for general research purposes through a request to I. Shai (email: [email protected]) and D. D. Wang (email: [email protected]) after publication. All of the source data for creating figures and extended data figures are available as supplementary information. Source data are provided with this paper.
Code availability
This study mainly relies on open-source bioinformatic tools described in detail in Methods . The analysis-specific programs are publicly available through https://github.com/DW-Group/T2D_Microbiome_Meta-analysis .
IDF Diabetes Atlas https://diabetesatlas.org/atlas/tenth-edition/ (2021).
American Diabetes Association Professional Practice Committee 2. Classification and diagnosis of diabetes: standards of medical care in diabetes – 2022. Diabetes Care 45 (Suppl. 1), S17–S38 (2022).
Article Google Scholar
Canfora, E. E., Meex, R. C. R., Venema, K. & Blaak, E. E. Gut microbial metabolites in obesity, NAFLD and T2DM. Nat. Rev. Endocrinol. 15 , 261–273 (2019).
Article CAS PubMed Google Scholar
Forslund, K. et al. Disentangling type 2 diabetes and metformin treatment signatures in the human gut microbiota. Nature 528 , 262–266 (2015).
Article CAS PubMed PubMed Central Google Scholar
Karlsson, F. H. et al. Gut metagenome in European women with normal, impaired and diabetic glucose control. Nature 498 , 99–103 (2013).
Qin, J. et al. A metagenome-wide association study of gut microbiota in type 2 diabetes. Nature 490 , 55–60 (2012).
Reitmeier, S. et al. Arrhythmic gut microbiome signatures predict risk of type 2 diabetes. Cell Host Microbe 28 , 258–272 (2020).
Sankaranarayanan, K. et al. Gut microbiome diversity among Cheyenne and Arapaho individuals from Western Oklahoma. Curr. Biol. 25 , 3161–3169 (2015).
Thingholm, L. B. et al. Obese individuals with and without type 2 diabetes show different gut microbial functional capacity and composition. Cell Host Microbe 26 , 252–264 (2019).
Wu, H. et al. The gut microbiota in prediabetes and diabetes: a population-based cross-sectional study. Cell Metab. 32 , 379–390 (2020).
Zhong, H. et al. Distinct gut metagenomics and metaproteomics signatures in prediabetics and treatment-naïve type 2 diabetics. EBioMedicine 47 , 373–383 (2019).
Article PubMed PubMed Central Google Scholar
Sonnenburg, J. L. & Bäckhed, F. Diet–microbiota interactions as moderators of human metabolism. Nature 535 , 56–64 (2016).
Pedersen, H. K. et al. Human gut microbes impact host serum metabolome and insulin sensitivity. Nature 535 , 376–381 (2016).
Dobrindt, U., Chowdary, M. G., Krumbholz, G. & Hacker, J. Genome dynamics and its impact on evolution of Escherichia coli . Med. Microbiol. Immunol. 199 , 145–154 (2010).
Van Rossum, T., Ferretti, P., Maistrenko, O. M. & Bork, P. Diversity within species: interpreting strains in microbiomes. Nat. Rev. Microbiol. 18 , 491–506 (2020).
Fromentin, S. et al. Microbiome and metabolome features of the cardiometabolic disease spectrum. Nat. Med. 28 , 303–314 (2022).
Yaskolka Meir, A. et al. Effect of green-Mediterranean diet on intrahepatic fat: the DIRECT PLUS randomised controlled trial. Gut 70 , 2085–2095 (2021).
Article PubMed Google Scholar
Pirzada, A. et al. Evolving science on cardiovascular disease among Hispanic/Latino adults. J. Am. Coll. Cardiol. 81 , 505–1520 (2023).
Mehta, R. S. et al. Stability of the human faecal microbiome in a cohort of adult men. Nat. Microbiol. 3 , 347–355 (2018).
Bao, Y. Origin, methods, and evolution of the Three Nurses' Health Studies. Am. J. Public Health. 105 , 1573–1581 (2016).
Beghini, F. et al. Integrating taxonomic, functional, and strain-level profiling of diverse microbial communities with bioBakery 3. Elife 10 , e65088 (2021).
Ma, S. et al. Population structure discovery in meta-analyzed microbial communities and inflammatory bowel disease using MMUPHin. Genome Biol. 23 , 208 (2022).
Mallick, H. et al. Multivariable association discovery in population-scale meta-omics studies. PLoS Comput. Biol. 17 , e1009442 (2021).
Thomas, A. M. et al. Metagenomic analysis of colorectal cancer datasets identifies cross-cohort microbial diagnostic signatures and a link with choline degradation. Nat. Med. 25 , 667–678 (2019).
Ruuskanen, M. O. et al. Gut microbiome composition is predictive of incident type 2 diabetes in a population cohort of 5,572 Finnish adults. Diabetes Care 45 , 811–818 (2022).
Atarashi, K. et al. Ectopic colonization of oral bacteria in the intestine drives T(H)1 cell induction and inflammation. Science 358 , 359–365 (2017).
Clooney, A. G. et al. Ranking microbiome variance in inflammatory bowel disease: a large longitudinal intercontinental study. Gut 70 , 499–510 (2021).
Cohen-Poradosu, R., McLoughlin, R. M., Lee, J. C. & Kasper, D. L. Bacteroides fragilis -stimulated interleukin-10 contains expanding disease. J. Infect. Dis. 204 , 363–371 (2011).
Garcia-Lopez, M. et al. Analysis of 1,000 type-strain genomes improves taxonomic classification of Bacteroidetes. Front. Microbiol. 10 , 2083 (2019).
Petersen, C. et al. T cell-mediated regulation of the microbiota protects against obesity. Science 365 , eaat9351 (2019).
Fung, T. C. et al. Intestinal serotonin and fluoxetine exposure modulate bacterial colonization in the gut. Nat. Microbiol. 4 , 2064–2073 (2019).
Riester, M. et al. Risk prediction for late-stage ovarian cancer by meta-analysis of 1,525 patient samples. J. Natl Cancer Inst. 106 , dju048 (2014).
Wu, H. et al. Metformin alters the gut microbiome of individuals with treatment-naïve type 2 diabetes, contributing to the therapeutic effects of the drug. Nat. Med. 23 , 850–858 (2017).
Forslund, S. K. et al. Combinatorial, additive and dose-dependent drug–microbiome associations. Nature 600 , 500–505 (2021).
Caspi, R. et al. The MetaCyc database of metabolic pathways and enzymes: a 2019 update. Nucleic Acids Res. 48 , D445–D453 (2020).
Anastasi, A., Knight, C. G. & Barrett, A. J. Characterization of the bacterial metalloendopeptidase pitrilysin by use of a continuous fluorescence assay. Biochem. J. 290 , 601–607 (1993).
Roden, M. & Shulman, G. I. The integrative biology of type 2 diabetes. Nature 576 , 51–60 (2019).
Vatanen, T. et al. Variation in microbiome LPS immunogenicity contributes to autoimmunity in humans. Cell 165 , 842–853 (2016).
Wang, D. D. et al. The gut microbiome modulates the protective association between a Mediterranean diet and cardiometabolic disease risk. Nat. Med. 27 , 333–343 (2021).
Wang, D. D. et al. The gut microbiome modifies the association between a Mediterranean diet and diabetes in USA Hispanic/ Latino population. J. Clin. Endocrinol. Metab. 107 , e924–e934 (2022).
Tett, A. et al. The Prevotella copri complex comprises four distinct clades underrepresented in westernized populations. Cell Host Microbe 26 , 666–679 (2019).
Truong, D. T., Tett, A., Pasolli, E., Huttenhower, C. & Segata, N. Microbial strain-level population structure and genetic diversity from metagenomes. Genome Res. 27 , 626–638 (2017).
Vangay, P. et al. US immigration westernizes the human gut microbiome. Cell 175 , 962–972 (2018).
Wang, T. J. et al. Metabolite profiles and the risk of developing diabetes. Nat. Med. 17 , 448–453 (2011).
Karcher, N. et al. Analysis of 1,321 Eubacterium rectale genomes from metagenomes uncovers complex phylogeographic population structure and subspecies functional adaptations. Genome Biol. 21 , 138 (2020).
Beghini, F. et al. Large-scale comparative metagenomics of Blastocystis , a common member of the human gut microbiome. ISME J. 11 , 2848–2863 (2017).
Hildebrand, F. et al. Dispersal strategies shape persistence and evolution of human gut bacteria. Cell Host Microbe 29 , 1167–1176 (2021).
Kaper, J. B., Nataro, J. P. & Mobley, H. L. Pathogenic Escherichia coli . Nat. Rev. Microbiol. 2 , 123–140 (2004).
Borodovich, T., Shkoporov, A. N., Ross, R. P. & Hill, C. Phage-mediated horizontal gene transfer and its implications for the human gut microbiome. Gastroenterol. Rep. 10 , goac012 (2022).
Bobay, L. M., Traverse, C. C. & Ochman, H. Impermanence of bacterial clones. Proc. Natl Acad. Sci. USA 112 , 8893–8900 (2015).
Navarro-Garcia, F. & Elias, W. P. Autotransporters and virulence of enteroaggregative E. coli . Gut Microbes 2 , 13–24 (2011).
Cani, P. D. et al. Metabolic endotoxemia initiates obesity and insulin resistance. Diabetes 56 , 1761–1772 (2007).
Vazquez-Lopez, J. & Navarro-Garcia, F. In silico analyses of core proteins and putative effector and immunity proteins for T6SS in enterohemorrhagic E. coli . Front. Cell. Infect. Microbiol. 10 , 195 (2020).
Ahmed, S. A. et al. Genomic comparison of Escherichia coli O104:H4 isolates from 2009 and 2011 reveals plasmid, and prophage heterogeneity, including Shiga toxin encoding phage stx2. PLoS One 7 , e48228 (2012).
Sun, H. et al. Regulation of flagellar motility and biosynthesis in enterohemorrhagic Escherichia coli O157:H7. Gut Microbes 14 , 2110822 (2022).
Chaban, B., Hughes, H. V. & Beeby, M. The flagellum in bacterial pathogens: for motility and a whole lot more. Semin. Cell Dev. Biol. 46 , 91–103 (2015).
Lux, R. & Shi, W. Chemotaxis-guided movements in bacteria. Crit. Rev. Oral Biol. Med. 15 , 207–220 (2004).
Ng, W. L. & Bassler, B. L. Bacterial quorum-sensing network architectures. Annu. Rev. Genet. 43 , 197–222 (2009).
Everard, A. et al. Cross-talk between Akkermansia muciniphila and intestinal epithelium controls diet-induced obesity. Proc. Natl Acad. Sci. USA 110 , 9066–9071 (2013).
Hillmann, B. et al. Evaluating the information content of shallow shotgun metagenomics. mSystems 3 , e00069-18 (2018).
IHMS Consortium. IHMS_SOP 03 V1: Standard Operating Procedure for Fecal Samples self-collection, laboratory analysis handled within 4 to 24 hours (4 hours < x ≤ 24 hours) , (International Human Microbiome Standards, 2015).
Salonen, A. et al. Comparative analysis of fecal DNA extraction methods with phylogenetic microarray: effective recovery of bacterial and archaeal DNA using mechanical cell lysis. J. Microbiol. Methods 81 , 127–134 (2010).
Courtois, S. et al. Recombinant environmental libraries provide access to microbial diversity for drug discovery from natural products. Appl. Environ. Microbiol. 69 , 49–55 (2003).
Deschasaux, M. et al. Depicting the composition of gut microbiota in a population with varied ethnic origins but shared geography. Nat. Med. 24 , 1526–1531 (2018).
Fang, C. et al. Assessment of the cPAS-based BGISEQ-500 platform for metagenomic sequencing. Gigascience 7 , 1–8 (2018).
Matthews, D. R. et al. Homeostasis model assessment: insulin resistance and beta-cell function from fasting plasma glucose and insulin concentrations in man. Diabetologia 28 , 412–419 (1985).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9 , 357–359 (2012).
Suzek, B. E., Huang, H., McGarvey, P., Mazumder, R. & Wu, C. H. UniRef: comprehensive and non-redundant UniProt reference clusters. Bioinformatics 23 , 1282–1288 (2007).
Buchfink, B., Xie, C. & Huson, D. H. Fast and sensitive protein alignment using DIAMOND. Nat. Methods 12 , 59–60 (2015).
Caspi, R. et al. The MetaCyc database of metabolic pathways and enzymes and the BioCyc collection of pathway/genome databases. Nucleic Acids Res. 44 , D471–D480 (2016).
Ye, Y. & Doak, T. G. A parsimony approach to biological pathway reconstruction/inference for genomes and metagenomes. PLoS Comput. Biol. 5 , e1000465 (2009).
Mirdita, M. et al. Uniclust databases of clustered and deeply annotated protein sequences and alignments. Nucleic Acids Res. 45 , D170–D176 (2017).
Johnson, W. E., Li, C. & Rabinovic, A. Adjusting batch effects in microarray expression data using empirical Bayes methods. Biostatistics 8 , 118–127 (2007).
Pasolli, E., Truong, D. T., Malik, F., Waldron, L. & Segata, N. Machine learning meta-analysis of large metagenomic datasets: tools and biological insights. PLoS Comput. Biol. 12 , e1004977 (2016).
Carpenter, B. et al. Stan: a probabilistic programming language. J. Stat. Softw. 76 , 1 (2017).
Paradis, E., Claude, J. & Strimmer, K. APE: analyses of phylogenetics and evolution in R language. Bioinformatics 20 , 289–290 (2004).
Vehtari, A., Gelman, A. & Gabry, J. Practical Bayesian model evaluation using leave-one-out cross-validation and WAIC. Stat. Comput. 27 , 1413–1432 (2017).
Zhou, X., Kao, M. C. & Wong, W. H. Transitive functional annotation by shortest-path analysis of gene expression data. Proc. Natl Acad. Sci. USA 99 , 12783–12788 (2002).
Download references
Acknowledgements
The authors thank K. Dennis for coordinating the collection and transfer of the data, and F. Bäckhed, K. Kristiansen, J. Li, H. Zhong and J. Qin for sharing their data and helping with the data transfer. The authors are indebted to the participants in the Health Professionals Follow-up Study (HPFS) and Nurses’ Health Study II (NHSII) for their continuing outstanding level of cooperation, and to the staff of the HPFS and NHSII for their valuable contributions. The authors also thank the staff and participants of the Hispanic Community Health Study/Study of Latinos (HCHS/SOL) for their important contributions; the DIRECT-PLUS Study participants for their valuable contributions; and A. Yaskolka-Meir, G. Tsaban, A. Kaplan, H. Zelica, I. Youngster, K. Tuohy and O. Koren for their contribution to the DIRECT-PLUS Study. This work is funded by the National Institute of Diabetes and Digestive and Kidney Diseases (NIDDK; R00 DK119412 and Boston Nutrition Obesity Research Center Pilot and Feasibility Program grant supported by P30 DK046200 to D.D.W.; R24 DK110499 to C.H.), National Institute of Nursing Research (R01 NR01999 to D.D.W.), National Institute on Aging (R01 AG077489 and RF1 AG083764 to D.D.W.) and National Cancer Institute (NCI; R35 CA253185 to A.T.C.). A.T.C. is an American Cancer Society Research Professor. F.W. is supported by the American Heart Association Postdoctoral Fellowship (Grant number: 897161 to F.W.). The HPFS is supported by research grants U01 CA167552 (to W.C.W.) and R01 HL035464 (to E.B.R.) from the National Institutes of Health (NIH). The Men’s Lifestyle Validation Study in HPFS was supported by U01 CA152904 (to M.J.S. and E.B.R.) from NCI. The fecal sample collection and metagenomic data sequencing in HPFS were supported by the STARR Cancer Consortium Award (I7-A714 to C.H.). NHSII was supported by U01 CA176726 from NIH and P01 CA055075 (to W.C.W.) from NCI. The fecal sample collection and metagenomic data sequencing in NHSII were supported by the R01 CA202704 (to A.T.C. and C.H.) from NCI. The HCHS/SOL is a collaborative study supported by contracts from the National Heart, Lung and Blood Institute (NHLBI) to the University of North Carolina (HHSN268201300001I/N01-HC-65233), University of Miami (HHSN268201300004I/N01-HC-65234), Albert Einstein College of Medicine (HHSN268201300002I/N01-HC-65235), University of Illinois at Chicago (HHSN268201300003I/N01-HC-65236 Northwestern University) and San Diego State University (HHSN268201300005I/N01-HC-65237). The following institutes, centers and/or offices have contributed to the HCHS/SOL through a transfer of funds to the NHLBI: National Institute on Minority Health and Health Disparities (NIMHD), National Institute on Deafness and Other Communication Disorders, National Institute of Dental and Craniofacial Research, NIDDK, National Institute of Neurological Disorders and Stroke, and NIH Institution-Office of Dietary Supplements. Additional funding for the ‘Gut Origins of Latino Diabetes’ ancillary study to HCHS/SOL was provided by R01 MD011389 (to R.C.K., R.D.B. and R.K.) from the NIMHD and the Life Course Methodology Core at Albert Einstein College of Medicine and the New York Regional Center for Diabetes Translation Research (P30 DK111022-8786 and P30 DK111022) through funds from NIDDK. Additional funding for this work was provided by R01 HL060712 (to F.B.H. and Q.Q.) from NHLBI. The DIRECT-PLUS Study was funded by grants from the Deutsche Forschungsgemeinschaft (DFG, German Research Foundation) Collaborative Research Center SFB1052 ‘Obesity Mechanisms’ (SFB-1052/B11 to I.S.); Israel Ministry of Health grant 87472511 (to I.S.); Israel Ministry of Science and Technology grant 3-13604 (to I.S.); California Walnuts Commission (to I.S.) and the CABALA_DIET&HEALTH Project, which received funding from the European Union’s Horizon 2020 Programme. The funding source had no role in the design and conduct of the study; collection, management, analysis and interpretation of the data; preparation, review, or approval of the manuscript; and the decision to submit the manuscript for publication. The content is solely the responsibility of the authors and does not necessarily represent the official views of the NIH. The computations in this paper were run in part on the FASRC Cannon cluster supported by the FAS Division of Science Research Computing Group at Harvard University.
Author information
These authors contributed equally: Zhendong Mei, Fenglei Wang.
Authors and Affiliations
Channing Division of Network Medicine, Department of Medicine, Brigham and Women’s Hospital and Harvard Medical School, Boston, MA, USA
Zhendong Mei, Danyue Dong, Yuxi Liu, Eric B. Rimm, Walter C. Willett, Frank B. Hu, Meir J. Stampfer & Dong D. Wang
Broad Institute of MIT and Harvard, Cambridge, MA, USA
Zhendong Mei, Fenglei Wang, Amrisha Bhosle, Danyue Dong, Raaj Mehta, Andrew Ghazi, Yancong Zhang, Yuxi Liu, Andrew T. Chan, Curtis Huttenhower & Dong D. Wang
Department of Nutrition, Harvard T.H. Chan School of Public Health, Boston, MA, USA
Fenglei Wang, Eric B. Rimm, Walter C. Willett, Frank B. Hu, Qibin Qi, Meir J. Stampfer, Iris Shai & Dong D. Wang
Department of Biostatistics, Harvard T.H. Chan School of Public Health, Boston, MA, USA
Amrisha Bhosle, Andrew Ghazi, Yancong Zhang & Curtis Huttenhower
Division of Gastroenterology, Massachusetts General Hospital and Harvard Medical School, Boston, MA, USA
Raaj Mehta & Andrew T. Chan
Clinical and Translational Epidemiology Unit, Massachusetts General Hospital and Harvard Medical School, Boston, MA, USA
Department of Chemistry and Chemical Biology, Harvard University, Cambridge, MA, USA
Department of Epidemiology, Harvard T.H. Chan School of Public Health, Boston, MA, USA
Yuxi Liu, Eric B. Rimm, Walter C. Willett, Frank B. Hu & Meir J. Stampfer
Department of Medicine, Hebrew University and Hadassah Medical Center, Jerusalem, Israel
Ehud Rinott
Department of Biostatistics, Vanderbilt University Medical Center, Nashville, TN, USA
Institute for Minority Health Research, University of Illinois Chicago, Chicago, IL, USA
Martha Daviglus
Center for Microbiome Innovation, Jacobs School of Engineering, University of California San Diego, La Jolla, CA, USA
Department of Pediatrics, School of Medicine, University of California San Diego, La Jolla, CA, USA
Department of Computer Science and Engineering, University of California San Diego, La Jolla, CA, USA
Department of Epidemiology and Population Health, Albert Einstein College of Medicine, Bronx, NY, USA
Qibin Qi, Robert D. Burk & Robert C. Kaplan
Department of Pediatrics, Albert Einstein College of Medicine, Bronx, NY, USA
Robert D. Burk
Department of Microbiology and Immunology, Albert Einstein College of Medicine, Bronx, NY, USA
Department of Obstetrics, Gynecology and Women’s Health, Albert Einstein College of Medicine, Bronx, NY, USA
Faculty of Health Sciences, The Health and Nutrition Innovative International Research Center, Ben-Gurion University of the Negev, Be′er Sheva, Israel
Public Health Sciences Division, Fred Hutchinson Cancer Research Center, Seattle, WA, USA
Robert C. Kaplan
Department of Immunology and Infectious Diseases, Harvard T.H. Chan School of Public Health, Boston, MA, USA
Curtis Huttenhower
Harvard Chan Microbiome in Public Health Center, Harvard T.H. Chan School of Public Health, Boston, MA, USA
You can also search for this author in PubMed Google Scholar
Contributions
Z.M., F.W., C.H. and D.D.W. conceptualized the study. Z.M. and F.W. performed the data analysis. Z.M., F.W., C.H. and D.D.W. drafted the paper. C.H. and D.D.W. supervised the study. E.B.R, M.D., W.C.W., R.K., F.B.H., Q.Q., A.T.C., R.D.B., M.J.S., E.R., I.S., R.C.K., C.H. and D.D.W. collected the sample and data, and obtained funding. Z.M., F.W., A.B., D.D., R.M., A.G., Y.Z., Y.L., E.R., S.M., E.B.R., M.D., W.C.W., R.K., F.B.H., Q.Q., A.T.C., R.D.B., M.J.S., I.S., R.C.K., C.H., and D.D.W. discussed the results, critically reviewed the text and approved the final paper.
Corresponding authors
Correspondence to Curtis Huttenhower or Dong D. Wang .
Ethics declarations
Competing interests.
C.H. is a member of the scientific advisory board for Zoe Nutrition, Empress Therapeutics, and Seres Therapeutics. All other authors have no competing interests.
Peer review
Peer review information.
Nature Medicine thanks the anonymous reviewers for their contribution to the peer review of this work. Primary Handling Editor: Sonia Muliyil, in collaboration with the Nature Medicine team.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended data fig. 1 workflow..
We adjusted for the study effect by adopting a conservative meta-analysis approach in the downstream analyses. Our analyses examined the overall microbial community structure, specific microbial taxonomic and functional features, strain-specific biochemical pathways, and within-species phylogeny and gene families in a cross-cohort meta-analysis framework. This figure was created with BioRender.com .
Extended Data Fig. 2 Principal coordinate analysis of all samples using species-level Bray–Curtis dissimilarity colored by cohorts before and after correcting batch and study effects.
R 2 values are calculated from permutational multivariate analysis of variance (PERMANOVA, n = 999 permutations) and indicate the variance attributable to study and batch effects.
Source data
Extended data fig. 3 comparisons in associations between microbial species and type 2 diabetes across different statistical models..
Meta-analyzed associations of individual microbial species with type 2 diabetes (T2D) phenotype from the ordinal ( a ) and binary ( b ) models. The ordinal model modeled the disease status as an ordinal variable (T2D, prediabetes, or controls) and used data from all the participants. The binary model modeled the disease status as a binary variable (T2D or controls) and used data from T2D patients and normoglycemic controls. The blue-to-red and purple-to-orange gradients represent the magnitude and direction of the associations as quantified by meta-analyzed beta coefficients from linear mixed models adjusted for age, sex, and body mass index (BMI) and further adjusted for metformin use in MaAsLin2. All the results were corrected for multiple hypothesis testing by controlling the false discovery rate (FDR) using the Benjamini–Hochberg method with a target rate of 0.10. All models included each participant’s identifier as random effects and simultaneously adjusted for covariables. ( c ) Comparisons in associations between microbial species and T2D between multivariate MaAsLin2 models with and without further adjustment for BMI and metformin use from the ordinal model. ( d ) Comparisons in associations between microbial species and T2D between multivariate MaAsLin2 models with and without further adjustment for BMI and metformin use from the binary model. Dots in the scatter plots in (c) and (d) represent meta-analyzed beta coefficients from linear mixed models adjusted for covariables in MaAsLin2. All the statistical tests were two-sided. A total of 8,117 metagenomes from 1,851 T2D patients, 2,770 individuals with prediabetes, and 2,277 normoglycemic controls were included in the analyses in (a), (b), (c), and (d). Abbreviations: BMI, body mass index; Con, control; metf, metformin use; insul, insulin use; T2D, type 2 diabetes.
Extended Data Fig. 4 Metformin has a direct impact on the gut microbiome composition and confounds the associations between microbial species and type 2 diabetes.
( a ) Distance-based redundancy analysis (dbRDA) based on species-level Bray–Curtis dissimilarity colored by type 2 diabetes (T2D) and metformin use. The centers of the boxplot show medians with boxes indicating their inter-quartile ranges (IQRs) and upper and lower whiskers indicating 1.5 times the IQR from above the upper quartile and below the lower quartile, respectively. ( b ) Meta-analyzed and cohort-specific associations of microbial species with metformin use among T2D patients. We defined microbial signatures of metformin as those significantly associated with metformin use in T2D cases only but not associated with T2D after further adjusting for metformin use in all participants. We also identified 4 species associated with both metformin use and T2D. The centers of the error bars represent the β coefficients of the associations, and the error bars represent their standard errors (SEs). ( c ) Our modeling approach effectively accounted for the potential confounding effect of metformin use, as evidenced by the high correlation between the beta coefficients of species–T2D associations obtained in the primary analysis and those calculated in a sensitivity analysis excluding T2D patients treated with metformin. The beta coefficients in (b) and (c) represent the associations quantified by linear mixed models, adjusting for age, sex, body mass index (BMI), and metformin use where appropriate, in MaAsLin2. All the results were corrected for multiple hypothesis testing by controlling the false discovery rate (FDR) using the Benjamini–Hochberg method with a target rate of 0.10. All the analyses in (a), (b), and (c) were based on 5,114 metagenomes from 1,851 T2D patients and 2,277 normoglycemic controls. The statistical tests in (a) and (b) were two-sided. Abbreviations: Con, control; metf, metformin use; T2D, type 2 diabetes.
Extended Data Fig. 5 Sensitivity analyses demonstrate that identified microbial features of type 2 diabetes are unlikely to reflect the duration or comorbidities of this disease.
( a ) Comparisons in associations between microbial species and T2D in one analysis that includes all study participants and the other that excludes individuals with prevalent T2D in the Hispanic Community Health Study/Study of Latinos. ( b ) Comparisons in associations between microbial species and T2D in one analysis that includes all study participants and the other analysis that excludes insulin-treated T2D patients. The dots represent the associations quantified by linear mixed models, adjusting for age, sex, body mass index, and metformin use in MaAsLin2. Abbreviation: T2D, type 2 diabetes.
Extended Data Fig. 6 Associations of microbial features with circulating metabolic and inflammation biomarkers.
( a ) Meta-analyzed associations of individual MetaCyc pathways with circulating biomarkers of metabolic risk. ( b ) Meta-analyzed associations of individual microbial enzymes with circulating biomarkers of metabolic risk. Only pathways and enzymes listed in Fig. 3 were analyzed and presented in this figure. The blue-to-red gradients represent the magnitude and direction of the associations as quantified by meta-analyzed beta coefficients from linear mixed models adjusted for age, sex, body mass index, and metformin use in MaAsLin2. All the results were corrected for multiple hypothesis testing by controlling the false discovery rate (FDR) using the Benjamini–Hochberg method with a target rate of 0.10. Abbreviations: BMI, body mass index; HbA1c, hemoglobin A1c; HDL-C, high-density lipoprotein cholesterol; hs-CRP, high-sensitivity C-reactive protein; HOMA-B, homeostasis model assessment of β-cell function; HOMA-IR, homeostasis model assessment of insulin resistance; LDL-C, low-density lipoprotein cholesterol; TG, triglyceride.
Extended Data Fig. 7 Prevotella copri ’s differential carriage of branched-chain amino acid biosynthesis function is explained by its discrete subclade structure.
( a ) Distribution of different P. copri subclades across geographic regions and studies. We applied MetaPhlAn taxonomic profiling based on P. copri subclade-specific marker genes to detect the presence of a subclade in metagenomes. ( b ) Comparisons in adjusted relative abundance of branched-chain amino acid (BCAA) biosynthesis pathways and enzyme encoded by P. copri subclades dominated by clade A versus other clades. The adjusted relative abundance of pathways and enzymes is estimated by anpan (ANalysis of microbial Phylogenies And geNes)’s pathway random effects models ( Methods ) with simultaneous adjustment for the abundance of P. copri subclades. The centers of the boxplot show medians of adjusted relative abundance with boxes indicating their inter-quartile ranges (IQRs) and upper and lower whiskers indicating 1.5 times the IQR from above the upper quartile and below the lower quartile, respectively. P -values were generated from two-sided t-tests based on the adjusted relative abundance. ( c ) Clade A-dominant P. copri strains in type 2 diabetes (T2D) patients were more likely to retain pathways and enzymes of branched-chain amino acid biosynthesis compared to clade A-dominant nonT2D controls. The blue and red lines, fitted by linear regression in participants with T2D and control participants separately, represent the associations between the log-transformed relative abundance of P. copri subclade and the log-transformed relative abundance of a given pathway or enzyme encoded by P. copri . The numeric values in the top left corner are posterior differences and 98% posterior intervals of differences in log-transformed pathway abundance between case–control status, as determined by mixed effects models anpan ( Methods ). This model allows us to identify microbial functions encoded by a P. copri subclade that are differentially abundant between T2D cases versus controls while controlling for its subclade-level abundance. All the analyses in (a), (b), and (c) were based on 5,114 metagenomes from 1,851 T2D patients and 2,277 normoglycemic controls.
Extended Data Fig. 8 Phylogenetic trees of select species show divergent associations between subclades and type 2 diabetes within each species.
The annotation bars represent metformin use (metf), study, body mass index (BMI), sex, age, and type 2 diabetes (T2D) status, respectively. The boxplots in the bottom represent the posterior mean of the phylogenetic effect of each phylogenetic tree leaf (metagenome) estimated by the phylogenetic generalized linear mixed models (PGLMMs) in anpan (ANalysis of microbial Phylogenies And geNes, see Methods ) with whiskers representing the 95% credible intervals of the posterior means. By applying PGLMMs, we compared two generalized linear mixed models with and without incorporating within-species phylogeny as a random effect ( Methods ). Both models were adjusted for age, sex, body mass index, metformin use, and study membership as fixed effects. We generated within-species phylogenetic trees by randomly splitting the edges based on the Euclidean similarity matrix derived from clustered sets of protein sequences (UniRef90 gene families) after dimension reduction by principal components analysis.
Extended Data Fig. 9 Gene set enrichment analysis of gene ontology terms for biological process.
The line plots show the running enrichment score for the gene ontology (GO) term as the analysis ‘walks down’ the ranked list. The vertical black lines on the X-axis show where members of the GO term appear in the ranked list of UniRef90 gene families.
Supplementary information
Supplementary information.
Supplementary Text and Supplementary Figs. 1 and 2
Reporting Summary
Supplementary Tables 1–13
Supplementary Data
Source Data for Supplementary Figs. 1 and 2
Source Data Fig. 1
Statistical Source Data
Source Data Fig. 2
Source data fig. 3, source data fig. 4, source data fig. 5, source data extended data fig. 2, source data extended data fig.3, source data extended data fig. 4, source data extended data fig. 5, source data extended data fig. 6, source data extended data fig. 7, source data extended data fig. 8, source data extended data fig. 9, rights and permissions.
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
Reprints and permissions
About this article
Cite this article.
Mei, Z., Wang, F., Bhosle, A. et al. Strain-specific gut microbial signatures in type 2 diabetes identified in a cross-cohort analysis of 8,117 metagenomes. Nat Med (2024). https://doi.org/10.1038/s41591-024-03067-7
Download citation
Received : 01 December 2023
Accepted : 14 May 2024
Published : 25 June 2024
DOI : https://doi.org/10.1038/s41591-024-03067-7
Share this article
Anyone you share the following link with will be able to read this content:
Sorry, a shareable link is not currently available for this article.
Provided by the Springer Nature SharedIt content-sharing initiative
Quick links
- Explore articles by subject
- Guide to authors
- Editorial policies
Sign up for the Nature Briefing: Microbiology newsletter — what matters in microbiology research, free to your inbox weekly.
Chemical Science
Unlocking the chemical environment of nitrogen in perovskite-type oxides †.

* Corresponding authors
a Institute of Multidisciplinary Research for Advanced Materials, Tohoku University, 2-1-1 Katahira, Aoba-ku, Sendai, Miyagi, Japan E-mail: [email protected] , [email protected]
b Advanced Institute for Materials Research (WPI-AIMR), Tohoku University, 2-1-1 Katahira, Aoba-ku, Sendai, Miyagi, Japan
c School of Environmental Science and Engineering, State Key Laboratory of Metal Matrix Composites, Shanghai Jiao Tong University, Shanghai, P. R. China
d Kyushu Synchrotron Light Research Center, 8-7 Yayoigaoka, Tosu, Saga, Japan
Nitrogen (N) doping of perovskite-type oxides is an effective method for enhancing their photocatalytic performance. Quantitative and qualitative analyses of the doped N species are essential for a deeper understanding of the catalytic activity enhancement mechanism. However, examining the N environment in perovskite-type oxides, particularly in the bulk, using conventional analytical techniques, such as X-ray photoelectron spectroscopy (XPS), is challenging. In this study, we propose a new analytical technique, advanced temperature-programmed desorption (TPD) up to 1600 °C, to complement the conventional methods. TPD can quantify all N species in bulk oxides. Moreover, it facilitates chemical speciation of N environments, such as substitutional and interstitial N species. This is verified by XPS, CHN elemental analysis, X-ray absorption spectroscopy, and in situ diffuse reflectance infrared Fourier-transform spectroscopy. This study demonstrates the feasibility of advanced TPD as a new analytical method that offers comprehensive information on the N species within N-doped oxide materials at the bulk level.

Supplementary files
- Supplementary information PDF (1805K)
Article information
Download Citation
Permissions.
Unlocking the chemical environment of nitrogen in perovskite-type oxides
S. Shimizu, T. Yoshii, G. Nishikawa, J. Wang, S. Yin, E. Kobayashi and H. Nishihara, Chem. Sci. , 2024, Advance Article , DOI: 10.1039/D4SC01850H
This article is licensed under a Creative Commons Attribution-NonCommercial 3.0 Unported Licence . You can use material from this article in other publications, without requesting further permission from the RSC, provided that the correct acknowledgement is given and it is not used for commercial purposes.
To request permission to reproduce material from this article in a commercial publication , please go to the Copyright Clearance Center request page .
If you are an author contributing to an RSC publication, you do not need to request permission provided correct acknowledgement is given.
If you are the author of this article, you do not need to request permission to reproduce figures and diagrams provided correct acknowledgement is given. If you want to reproduce the whole article in a third-party commercial publication (excluding your thesis/dissertation for which permission is not required) please go to the Copyright Clearance Center request page .
Read more about how to correctly acknowledge RSC content .
Social activity
Search articles by author.
This article has not yet been cited.
Advertisements

IMAGES
VIDEO
COMMENTS
Step 2: Methods. Next, indicate the research methods that you used to answer your question. This part should be a straightforward description of what you did in one or two sentences. It is usually written in the past simple tense, as it refers to completed actions.
An abstract is a brief summary of a completed research/innovation project. What should an abstract include? An abstract should include the following components: 1. Background: What is the motivation behind the research? Provide a short paragraph that details the background information. 2. Objectives: What problem are you attempting to solve ...
1. Write in Active Voice. First, use active voice when writing an abstract for your research proposal. However, this doesn't mean you should avoid passive voice in entirety. If you find that some sentences can't make sense unless with passive sentence construction, feel free to bend this rule somewhat.
5. How to Format an Abstract. Most abstracts use the same formatting rules, which help the reader identify the abstract so they know where to look for it. Here's a list of formatting guidelines for writing an abstract: Stick to one paragraph. Use block formatting with no indentation at the beginning.
Definition and Purpose of Abstracts An abstract is a short summary of your (published or unpublished) research paper, usually about a paragraph (c. 6-7 sentences, 150-250 words) long. A well-written abstract serves multiple purposes: an abstract lets readers get the gist or essence of your paper or article quickly, in order to decide whether to….
You can, however, write a draft at the beginning of your research and add in any gaps later. If you find abstract writing a herculean task, here are the few tips to help you with it: 1. Always develop a framework to support your abstract. Before writing, ensure you create a clear outline for your abstract.
An APA abstract is a brief, comprehensive summary of the contents of an article, research paper, dissertation, or report. It is written in accordance with the guidelines of the American Psychological Association (APA), which is a widely used format in social and behavioral sciences.
Follow these five steps to format your abstract in APA Style: Insert a running head (for a professional paper—not needed for a student paper) and page number. Set page margins to 1 inch (2.54 cm). Write "Abstract" (bold and centered) at the top of the page. Place the contents of your abstract on the next line.
Include 5 to 10 important words or short phrases central to your research in both the abstract and the keywords section. For example, if you are writing a paper on the prevalence of obesity among lower classes that crosses international boundaries, you should include terms like "obesity," "prevalence," "international," "lower ...
You will almost always have to include an abstract when: Completing a thesis or dissertation. Submitting a research paper to an academic journal. Writing a book proposal. Applying for research grants. It's easiest to write your abstract last, because it's a summary of the work you've already done.
Informative Abstract Example 1. Emotional intelligence (EQ) has been correlated with leadership effectiveness in organizations. Using a mixed-methods approach, this study assesses the importance of emotional intelligence on academic performance at the high school level. The Emotional Intelligence rating scale was used, as well as semi ...
Write your paper first, then create the abstract as a summary. Check the journal requirements before you write your abstract, eg. required subheadings. Include keywords or phrases to help readers search for your work in indexing databases like PubMed or Google Scholar. Double and triple check your abstract for spelling and grammar errors.
Research proposal examples. Writing a research proposal can be quite challenging, but a good starting point could be to look at some examples. We've included a few for you below. Example research proposal #1: "A Conceptual Framework for Scheduling Constraint Management".
HIV virus = Human Immunodeficiency Virus; AIDS syndrome = Acquired Immunodeficiency Syndrome. 2. Avoid Useless and Emotional Intensifiers. Really, very, quite, extremely, severely, clearly, certainly, essentially, actually: The preliminary results clearly show that the protein was absent in the fraction.
Abstracts provide a summary and preview of an academic work, such an article, research proposal, or conference presentation. Abstracts are the first part of an article that readers will see: They set expectations and help readers understand what will come next. All abstracts used in this handout are from published articles from biology ...
There are six steps to writing a standard abstract. (1) Begin with a broad statement about your topic. Then, (2) state the problem or knowledge gap related to this topic that your study explores. After that, (3) describe what specific aspect of this problem you investigated, and (4) briefly explain how you went about doing this.
An abstract summarizes, usually in one paragraph of 300 words or less, the major aspects of the entire paper in a prescribed sequence that includes: 1) the overall purpose of the study and the research problem(s) you investigated; 2) the basic design of the study; 3) major findings or trends found as a result of your analysis; and, 4) a brief summary of your interpretations and conclusions.
Set a 1-inch (2.54 centimeter) margin on all sides. The running head should be aligned to the left at the top of the page. The abstract should be on the second page of the paper (the first one is reserved for the title). Avoid indentations, unless you must include a keywords section at the end of the abstract.
problem under investigation or research question(s) • clearly stated hypothesis or hypotheses • methods used (including brief descriptions of the study design, sample, and sample size) • study results • • implications (i.e., why this study is important, applications of the results or findings) Abstract Format • recommended fonts: 11 ...
If you are writing an abstract as a proposal for your research—in other words, as a request for permission to write a paper—the abstract serves to predict the kind of paper you hope to write. ... The abstract should begin with a clear sense of the research question you have framed. Often writers set this up as a problem: "Although some ...
Helpful tips when writing an abstract: • Reread your article or proposal with the goal of abstracting in mind. o Look specifically for these main parts of the article or proposal: purpose, methods, scope, results, conclusions and recommendations. o Use the headings and table of contents as a guide to writing your abstract.
Follow Proper Formatting. First person point of view -- "I" and "my" -- are usually acceptable in APA proposals, but you should double check your field's style guide. After finishing a draft, revise your abstract to create concise language, keeping the abstract to a maximum of 250 words. Find examples of acceptable abstracts from your field and ...
Example 1. Reprocessing used nuclear fuel (UNF) is crucial to the completion of a closed fuel cycle and would reduce the volume of waste produced during nuclear power production. Pyroprocessing is a promising reprocessing technique as it offers pure forms of product recovery. A limiting issue with pyroprocessing, however, is the inability to ...
than 10,000 people and a nationally representative sample of CWSs and NTNCWSs serving 10,000 or fewer people. UCMR 5, published in 2021, includes samplingfor 29 PFAS (including the six PFAS required ... and the USDA Agricultural Research Service's Sustaining The Earth's Watersheds—Agricultural Research Database System (STEWARDS) (https ...
The sample processing and shotgun metagenomic sequencing were performed at Alkek Center for Metagenomics and Microbiome Research, Baylor College of Medicine, Houston, TX, United States.
a Institute of Multidisciplinary Research for Advanced Materials, Tohoku University, 2-1-1 Katahira, Aoba-ku, Sendai, Miyagi, Japan ... Abstract. Nitrogen (N) doping of perovskite-type oxides is an effective method for enhancing their photocatalytic performance. Quantitative and qualitative analyses of the doped N species are essential for a ...
Research Note Framing disinformation through legislation: Evidence from policy proposals in Brazil This article analyzes 62 bills introduced in the Brazilian Chamber of Deputies between 2019-2022 to understand how legislators frame disinformation into different problems and their respective solutions.